Showing Spotlights 1513 - 1520 of 2777 in category All (newest first):
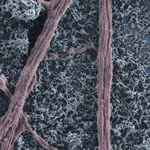 Carbon nanotubes, like the nervous cells of our brain, are excellent electrical signal conductors and can form intimate mechanical contacts with cellular membranes, thereby establishing a functional link to neuronal structures. There is a growing body of research on using nanomaterials in neural engineering. Now, researchers have, for the first time, explored the impact of carbon nanotube scaffolds on multilayered neuronal networks. Up to now, all known effects of carbon nanotubes on neurons - namely their reported ability to potentiate neuronal signaling and synapses - have been described in bi-dimensional cultured networks where nanotube/neuron hybrids were developed on a monolayer of dissociated brain cells.
Carbon nanotubes, like the nervous cells of our brain, are excellent electrical signal conductors and can form intimate mechanical contacts with cellular membranes, thereby establishing a functional link to neuronal structures. There is a growing body of research on using nanomaterials in neural engineering. Now, researchers have, for the first time, explored the impact of carbon nanotube scaffolds on multilayered neuronal networks. Up to now, all known effects of carbon nanotubes on neurons - namely their reported ability to potentiate neuronal signaling and synapses - have been described in bi-dimensional cultured networks where nanotube/neuron hybrids were developed on a monolayer of dissociated brain cells.
Mar 2nd, 2012
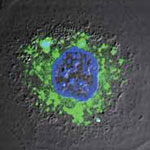 Carbon nanotubes (CNTs) offer a number of advantages for delivering drugs to specific locations inside the body which suggest that they may provide an improved result over nanoparticles. They have a larger inner volume which allows more drug molecules to be encapsulated, and this volume is more easily accessible because the end caps can be easily removed, and they have distinct inner and outer surfaces for functionalization. Recent research has shown the ability of CNTs to carry a variety of molecules such as drugs, DNA, proteins, peptides, targeting ligands etc. into cells - which makes them suitable candidates for targeted delivery applications. Researchers have now developed a unique two-dye labeling method to directly track the release process of a anti-cancer drug from carbon nanotube carriers in living cells.
Carbon nanotubes (CNTs) offer a number of advantages for delivering drugs to specific locations inside the body which suggest that they may provide an improved result over nanoparticles. They have a larger inner volume which allows more drug molecules to be encapsulated, and this volume is more easily accessible because the end caps can be easily removed, and they have distinct inner and outer surfaces for functionalization. Recent research has shown the ability of CNTs to carry a variety of molecules such as drugs, DNA, proteins, peptides, targeting ligands etc. into cells - which makes them suitable candidates for targeted delivery applications. Researchers have now developed a unique two-dye labeling method to directly track the release process of a anti-cancer drug from carbon nanotube carriers in living cells.
Mar 1st, 2012
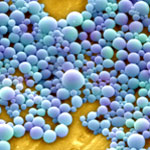 A group of experts from the chemical industry and various research laboratories in Germany have published a report on the current status of risk research on nanotechnology materials and applications. The report - 10 Years of Research: Risk Assessment, Human and Environmental Toxicology of Nanomaterials - provides an overview of the current state of risk assessment and toxicological research into nanomaterials. It also lists and summarizes the national and European projects on toxicology on various nanomaterials. In their report, the working group "Responsible Production and Use of Nanomaterials" has drawn up a list of topics and priorities which need to be addressed; activities and projects which have already been carried out; are currently on-going; or are still at the planning stage. The main focus of our considerations is on Germany, with a wider outlook on papers and results at European level.
A group of experts from the chemical industry and various research laboratories in Germany have published a report on the current status of risk research on nanotechnology materials and applications. The report - 10 Years of Research: Risk Assessment, Human and Environmental Toxicology of Nanomaterials - provides an overview of the current state of risk assessment and toxicological research into nanomaterials. It also lists and summarizes the national and European projects on toxicology on various nanomaterials. In their report, the working group "Responsible Production and Use of Nanomaterials" has drawn up a list of topics and priorities which need to be addressed; activities and projects which have already been carried out; are currently on-going; or are still at the planning stage. The main focus of our considerations is on Germany, with a wider outlook on papers and results at European level.
Feb 28th, 2012
 The significant research interest in the engineering of photovoltaic (PV) structures at the nanoscale is directed toward enabling reductions in PV module fabrication and installation costs as well as improving cell power conversion efficiency. With the emergence of a multitude of nanostructured photovoltaic device architectures, the question has arisen of where both the practical and the fundamental limits of performance reside in these new systems. A particular advantage of nanostructured materials is the tunability of their optical and electronic properties, which enables improved PV power conversion efficiencies by implementing strategies for reducing thermal losses. A recent review article addresses the limits to the performance of molecular, organic, polymeric, dye-sensitized, and colloidal quantum dot-based solar cells.
The significant research interest in the engineering of photovoltaic (PV) structures at the nanoscale is directed toward enabling reductions in PV module fabrication and installation costs as well as improving cell power conversion efficiency. With the emergence of a multitude of nanostructured photovoltaic device architectures, the question has arisen of where both the practical and the fundamental limits of performance reside in these new systems. A particular advantage of nanostructured materials is the tunability of their optical and electronic properties, which enables improved PV power conversion efficiencies by implementing strategies for reducing thermal losses. A recent review article addresses the limits to the performance of molecular, organic, polymeric, dye-sensitized, and colloidal quantum dot-based solar cells.
Feb 23rd, 2012
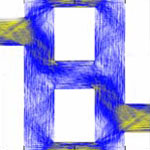 Researchers have demonstrated that electrons in nanoscale networks can behave like car drivers in congested cities. Traffic planners are sometimes faced with a rather counter-intuitive observation - adding a new road to a congested road network can lead to a deterioration of the overall traffic situation, i.e. longer trip times for individual road users. Or, in reverse, blocking certain streets in a complex road network can counter-intuitively reduce congestion. This has become known as the Braess paradox. Researchers have now applied the concept of the Braess paradox to the quantum world. By combining quantum simulations of a model system and scanning-probe experiments, they have shown that an analogue of the Braess paradox can occur in mesoscopic semiconductor networks, where electron transport is governed by quantum mechanics. The paradox manifests itself by an increase of the conductance network when one arm in the network is partially blocked in a controlled manner.
Researchers have demonstrated that electrons in nanoscale networks can behave like car drivers in congested cities. Traffic planners are sometimes faced with a rather counter-intuitive observation - adding a new road to a congested road network can lead to a deterioration of the overall traffic situation, i.e. longer trip times for individual road users. Or, in reverse, blocking certain streets in a complex road network can counter-intuitively reduce congestion. This has become known as the Braess paradox. Researchers have now applied the concept of the Braess paradox to the quantum world. By combining quantum simulations of a model system and scanning-probe experiments, they have shown that an analogue of the Braess paradox can occur in mesoscopic semiconductor networks, where electron transport is governed by quantum mechanics. The paradox manifests itself by an increase of the conductance network when one arm in the network is partially blocked in a controlled manner.
Feb 21st, 2012
 Graphene is an excellent conductor of heat and might be an ideal material for thermal management in nanoelectronics. In field-effect transistors or interconnects most of the heat propagates directly below the graphene channel in the direction of the heat sink, that is, the bottom of a silicon wafer. For this reason, the highly thermally resistive silicon layers act as a thermal bottleneck, preventing the full utilization of graphene's excellent intrinsic properties. The breakdown current density in typical graphene devices is a hundred times larger than the fundamental limit in metals, which is set by electromigration. Researchers wanted to see if we can push the breakdown current density in graphene even further by better removal of dissipated heat. They managed to do it with the help of high-quality synthetic diamond. The graphene transistors or interconnects on synthetic diamond can sustain current densities which are a thousand times larger than in metals.
Graphene is an excellent conductor of heat and might be an ideal material for thermal management in nanoelectronics. In field-effect transistors or interconnects most of the heat propagates directly below the graphene channel in the direction of the heat sink, that is, the bottom of a silicon wafer. For this reason, the highly thermally resistive silicon layers act as a thermal bottleneck, preventing the full utilization of graphene's excellent intrinsic properties. The breakdown current density in typical graphene devices is a hundred times larger than the fundamental limit in metals, which is set by electromigration. Researchers wanted to see if we can push the breakdown current density in graphene even further by better removal of dissipated heat. They managed to do it with the help of high-quality synthetic diamond. The graphene transistors or interconnects on synthetic diamond can sustain current densities which are a thousand times larger than in metals.
Feb 20th, 2012
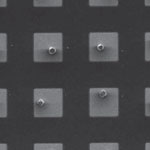 Metal-organic frameworks (MOFs) are well-ordered, lattice-like crystals. The nodes of the lattices are metals which are connected by organic molecules. Their controlled nanometer-sized pores provide MOFs with the potential to be used in next generation gas storage, gas separation and sensors. With their special structure and large surface area, MOFs open up new opportunities for alternative systems for gas and energy storage (e.g. carbon dioxide and hydrogen storage), in catalysis, chemical sensing, as nanoreactors, and in drug delivery, making them hugely interesting for both university research and industry. Researchers have now achieved the first microfluidic method for patterning MOF crystals. As such, high-throughput single crystal patterning is achieved with an unprecedented degree of flexibility.
Metal-organic frameworks (MOFs) are well-ordered, lattice-like crystals. The nodes of the lattices are metals which are connected by organic molecules. Their controlled nanometer-sized pores provide MOFs with the potential to be used in next generation gas storage, gas separation and sensors. With their special structure and large surface area, MOFs open up new opportunities for alternative systems for gas and energy storage (e.g. carbon dioxide and hydrogen storage), in catalysis, chemical sensing, as nanoreactors, and in drug delivery, making them hugely interesting for both university research and industry. Researchers have now achieved the first microfluidic method for patterning MOF crystals. As such, high-throughput single crystal patterning is achieved with an unprecedented degree of flexibility.
Feb 16th, 2012
 Following up on our recent Nanowerk Spotlight on nanofoods, new research shows that consumers could be exposed to nanoparticles in food by a much larger degree than has been expected so far. For a modern consumer it is hard to avoid titanium dioxide (TiO2) - a widely used additive in food, personal care and other household products. Approximately 7 million tons of bulk TiO2 are produced annually and used as white pigment in order to provide whiteness and opacity to foods and other products. Many applications of titanium dioxide would benefit from smaller primary particle sizes, and we can expect the percentage of TiO2 that is produced in or near the nano range to increase.
Following up on our recent Nanowerk Spotlight on nanofoods, new research shows that consumers could be exposed to nanoparticles in food by a much larger degree than has been expected so far. For a modern consumer it is hard to avoid titanium dioxide (TiO2) - a widely used additive in food, personal care and other household products. Approximately 7 million tons of bulk TiO2 are produced annually and used as white pigment in order to provide whiteness and opacity to foods and other products. Many applications of titanium dioxide would benefit from smaller primary particle sizes, and we can expect the percentage of TiO2 that is produced in or near the nano range to increase.
Feb 15th, 2012
 Carbon nanotubes, like the nervous cells of our brain, are excellent electrical signal conductors and can form intimate mechanical contacts with cellular membranes, thereby establishing a functional link to neuronal structures. There is a growing body of research on using nanomaterials in neural engineering. Now, researchers have, for the first time, explored the impact of carbon nanotube scaffolds on multilayered neuronal networks. Up to now, all known effects of carbon nanotubes on neurons - namely their reported ability to potentiate neuronal signaling and synapses - have been described in bi-dimensional cultured networks where nanotube/neuron hybrids were developed on a monolayer of dissociated brain cells.
Carbon nanotubes, like the nervous cells of our brain, are excellent electrical signal conductors and can form intimate mechanical contacts with cellular membranes, thereby establishing a functional link to neuronal structures. There is a growing body of research on using nanomaterials in neural engineering. Now, researchers have, for the first time, explored the impact of carbon nanotube scaffolds on multilayered neuronal networks. Up to now, all known effects of carbon nanotubes on neurons - namely their reported ability to potentiate neuronal signaling and synapses - have been described in bi-dimensional cultured networks where nanotube/neuron hybrids were developed on a monolayer of dissociated brain cells.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





