Showing Spotlights 49 - 56 of 77 in category All (newest first):
 Molecular-size motors have evolved in nature, where they are used in virtually every important biological process. In contrast, the development of synthetic nanomotors that mimic the function of these amazing natural systems and that could be used in man-made nanodevices is in its infancy. Building nanoscale motors is not just an exercise in scaling down the design of a macroworld engine to nanoscale dimensions. In addition to organic molecules, scientists increasingly are looking to DNA as a very promising way to fabricate nanomotors. The concept of a single DNA molecule nanomotor was already introduced in early 2002. However, this and subsequent designs require addition and removal of fuel and waste strands for motor function, although some artificial nanomotors can utilize alternative energy sources, including hydrolysis of the DNA backbone and ATP. Researchers at the University of Florida have now designed a photoswitchable single-molecule DNA nanomotor. It is the first fully reversible single-molecule DNA nanomachine driven by photons without any additional DNA strands as fuel.
Molecular-size motors have evolved in nature, where they are used in virtually every important biological process. In contrast, the development of synthetic nanomotors that mimic the function of these amazing natural systems and that could be used in man-made nanodevices is in its infancy. Building nanoscale motors is not just an exercise in scaling down the design of a macroworld engine to nanoscale dimensions. In addition to organic molecules, scientists increasingly are looking to DNA as a very promising way to fabricate nanomotors. The concept of a single DNA molecule nanomotor was already introduced in early 2002. However, this and subsequent designs require addition and removal of fuel and waste strands for motor function, although some artificial nanomotors can utilize alternative energy sources, including hydrolysis of the DNA backbone and ATP. Researchers at the University of Florida have now designed a photoswitchable single-molecule DNA nanomotor. It is the first fully reversible single-molecule DNA nanomachine driven by photons without any additional DNA strands as fuel.
Jun 10th, 2009
 Just a few days ago we covered the exciting and quickly developing world of nanotechnology machinery, specifically nanomotors. In this previous Nanowerk Spotlight we focused on one approach to nanomotors, which is copying nature's catalytic biomotors. Today we will look at an example of mechanical approach that works with carbon nanotubes (CNTs). Some experimental work concerning CNT motor system has already been reported, but new work coming out of Japan could very well be the smallest motor so far. Researchers in Japan investigated the linear and rotary motions of a CNT capsule at room temperature when it is sealed by other CNTs in a hollow space of a host CNT. It is the first observation of linear motion of CNT capsules. Such a system can be obtained by simply heating C60 peapods, and its size is comparable or much smaller than well-known protein-based molecular motors in the bioengineering field.
Just a few days ago we covered the exciting and quickly developing world of nanotechnology machinery, specifically nanomotors. In this previous Nanowerk Spotlight we focused on one approach to nanomotors, which is copying nature's catalytic biomotors. Today we will look at an example of mechanical approach that works with carbon nanotubes (CNTs). Some experimental work concerning CNT motor system has already been reported, but new work coming out of Japan could very well be the smallest motor so far. Researchers in Japan investigated the linear and rotary motions of a CNT capsule at room temperature when it is sealed by other CNTs in a hollow space of a host CNT. It is the first observation of linear motion of CNT capsules. Such a system can be obtained by simply heating C60 peapods, and its size is comparable or much smaller than well-known protein-based molecular motors in the bioengineering field.
Feb 17th, 2009
 While the concept of a 'machine' can be extended to the nano-world, these nanomachines can not be built by just further miniaturizing machine blueprints from the macro-world. On the nanoscale, the nanomachine components would be atomic or molecular structures each designed to perform a specific task which, all taken together, would result in a complex function. The problem is that functional nanomachinery will need to take into account the quantum effects that dominate the behavior of matter at the nanoscale, affecting the optical, electrical and magnetic behavior of materials. An alternative approach to miniaturizing machines down to the nanoscale is to borrow from the highly successful design shop of Mother Nature. Until a few years ago, catalytic micro- and nanomotors have been more or less unknown outside biology. With the explosive growth in nanotechnology, however, catalyzed nanoscale motion has become a heavily researched phenomenon. Scientists find that there is much to be learned from nature's motor systems for the development of artificial nanoscale machinery. Joseph Wang, who runs the Laboratory for Nanobioelectronics at the Department of NanoEngineering at UC San Diego, has just published a review where he addresses the question: Can Man-Made Nanomachines Compete with Nature Biomotors?
While the concept of a 'machine' can be extended to the nano-world, these nanomachines can not be built by just further miniaturizing machine blueprints from the macro-world. On the nanoscale, the nanomachine components would be atomic or molecular structures each designed to perform a specific task which, all taken together, would result in a complex function. The problem is that functional nanomachinery will need to take into account the quantum effects that dominate the behavior of matter at the nanoscale, affecting the optical, electrical and magnetic behavior of materials. An alternative approach to miniaturizing machines down to the nanoscale is to borrow from the highly successful design shop of Mother Nature. Until a few years ago, catalytic micro- and nanomotors have been more or less unknown outside biology. With the explosive growth in nanotechnology, however, catalyzed nanoscale motion has become a heavily researched phenomenon. Scientists find that there is much to be learned from nature's motor systems for the development of artificial nanoscale machinery. Joseph Wang, who runs the Laboratory for Nanobioelectronics at the Department of NanoEngineering at UC San Diego, has just published a review where he addresses the question: Can Man-Made Nanomachines Compete with Nature Biomotors?
Feb 4th, 2009
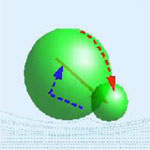 Advances in micro- and nanoscale engineering have led to various mobile devices that either can move on solids or swim in fluids. Researchers are applying various strategies to designing nanoscale propulsion systems by either using or copying biological systems such as the flagellar motors of bacteria or by employing various chemical reactions. Many of these approaches are fairly complex and not necessarily suited for large-scale deployment in practical applications. Scientists have theorized about simpler designs for mechanical swimmers that avoid the complexities of biological mechanisms and use very few degrees of freedom.
Researchers in Spain have now demonstrated the experimental realization of a simple device made by microscopic colloidal particles which can be externally controlled and propelled at low Reynolds number condition, i.e. when the viscosity of the fluid dominates over the inertia of the object. This is the same condition that governs the motion of bacteria such as E. Coli or other micro- or nanoscale objects that move in a fluid.
Advances in micro- and nanoscale engineering have led to various mobile devices that either can move on solids or swim in fluids. Researchers are applying various strategies to designing nanoscale propulsion systems by either using or copying biological systems such as the flagellar motors of bacteria or by employing various chemical reactions. Many of these approaches are fairly complex and not necessarily suited for large-scale deployment in practical applications. Scientists have theorized about simpler designs for mechanical swimmers that avoid the complexities of biological mechanisms and use very few degrees of freedom.
Researchers in Spain have now demonstrated the experimental realization of a simple device made by microscopic colloidal particles which can be externally controlled and propelled at low Reynolds number condition, i.e. when the viscosity of the fluid dominates over the inertia of the object. This is the same condition that governs the motion of bacteria such as E. Coli or other micro- or nanoscale objects that move in a fluid.
Dec 1st, 2008
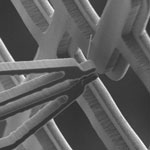 Nanotechnology researchers are actively working on the beginnings of various nanorobotic systems that one day could lead to automated, assembly-line type nanofabrication processes. Last year we reported on a nanogripper, a kind of a robotic 'hand' some ten thousand times smaller than a human hand. This 'pick-and-place' device used a silicon gripper which was controlled by a nanorobotic arm and was capable of picking up a carbon nanofiber and fix it onto the tip of an atomic force microscope cantilever. One of the problems that is vexing researchers is that the nanoscale miniaturization of these grippers comes at the cost of reduced strength - the smaller the gripper, the weaker it becomes. Therefore what is needed is a gripper design that is strong enough yet sufficient flexible and small to handle tough materials like carbon nanotubes.
Nanotechnology researchers are actively working on the beginnings of various nanorobotic systems that one day could lead to automated, assembly-line type nanofabrication processes. Last year we reported on a nanogripper, a kind of a robotic 'hand' some ten thousand times smaller than a human hand. This 'pick-and-place' device used a silicon gripper which was controlled by a nanorobotic arm and was capable of picking up a carbon nanofiber and fix it onto the tip of an atomic force microscope cantilever. One of the problems that is vexing researchers is that the nanoscale miniaturization of these grippers comes at the cost of reduced strength - the smaller the gripper, the weaker it becomes. Therefore what is needed is a gripper design that is strong enough yet sufficient flexible and small to handle tough materials like carbon nanotubes.
Nov 26th, 2008
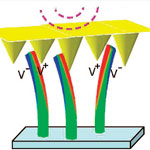 A major issue for using self-contained nanodevices, for instance as implantable nanosensors or environmental monitoring devices, is the question of how to power these tiny machines with an independent power source. Options include nanobatteries and nanogenerators that harvest energy from their environment. By converting mechanical energy from body movement, engine vibrations, or water or air flow into electricity, these nanoscale power plants could make possible a new class of self-powered medical devices, sensors and portable electronics. Probably the leading team that is driving forward the work on nanogenerators for converting mechanical energy into electricity is Zhong Lin Wang's group at Georgia Tech. Wang's team has now designed and demonstrated an innovative approach to fabricating a nanogenerator by integrating nanowires and pyramid-shaped nanobrushes into a multilayer power generator. By demonstrating an effective way for raising the output current, voltage and power, this work provides the technological platform for scaling up the performance of nanogenerators to a level that some day might be able to independently power devices like pacemakers or iPods.
A major issue for using self-contained nanodevices, for instance as implantable nanosensors or environmental monitoring devices, is the question of how to power these tiny machines with an independent power source. Options include nanobatteries and nanogenerators that harvest energy from their environment. By converting mechanical energy from body movement, engine vibrations, or water or air flow into electricity, these nanoscale power plants could make possible a new class of self-powered medical devices, sensors and portable electronics. Probably the leading team that is driving forward the work on nanogenerators for converting mechanical energy into electricity is Zhong Lin Wang's group at Georgia Tech. Wang's team has now designed and demonstrated an innovative approach to fabricating a nanogenerator by integrating nanowires and pyramid-shaped nanobrushes into a multilayer power generator. By demonstrating an effective way for raising the output current, voltage and power, this work provides the technological platform for scaling up the performance of nanogenerators to a level that some day might be able to independently power devices like pacemakers or iPods.
Oct 27th, 2008
 The concept of a 'machine' - a mechanical or electrical device that transmits or modifies energy to perform a certain task - can be extended to the nano-world. On the nanoscale, the nanomachine components would be atomic or molecular structures each designed to perform a specific task which, all taken together, would result in a complex function. However, these nanomachines can not be built by just further miniaturizing machine blueprints from the macro-world. Functional nanomachinery will need to take into account the quantum effects that dominate the behavior of matter at the nanoscale, affecting the optical, electrical and magnetic behavior of materials. Scientists in The Netherlands are now reporting how an atomic scale mechanical device consisting of two moving parts, each composed of only two atoms can be controlled by an external electrical signal, while being stable and providing a variety of functional modes. They jokingly refer to it as playing atomic pinball, since the two moving parts resemble the flippers in a pinball machine - unfortunately they haven't got a ball yet to play with.
The concept of a 'machine' - a mechanical or electrical device that transmits or modifies energy to perform a certain task - can be extended to the nano-world. On the nanoscale, the nanomachine components would be atomic or molecular structures each designed to perform a specific task which, all taken together, would result in a complex function. However, these nanomachines can not be built by just further miniaturizing machine blueprints from the macro-world. Functional nanomachinery will need to take into account the quantum effects that dominate the behavior of matter at the nanoscale, affecting the optical, electrical and magnetic behavior of materials. Scientists in The Netherlands are now reporting how an atomic scale mechanical device consisting of two moving parts, each composed of only two atoms can be controlled by an external electrical signal, while being stable and providing a variety of functional modes. They jokingly refer to it as playing atomic pinball, since the two moving parts resemble the flippers in a pinball machine - unfortunately they haven't got a ball yet to play with.
Sep 23rd, 2008
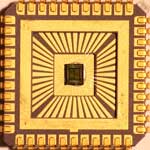 For nanoelectronics applications like single-electron devices to become practical, everyday items, they need to move from the highly individual and customized fabrication process typically found in laboratories to an automated, high-throughput and industrial-scale production environment. The reason this hasn't happened yet is that the various nanoscale pattern definition techniques used today - such as e-beam lithography, mechanically controllable break junctions, electromigration, electrodeposition, nanoscale oxidation, and scanning tunneling microscopy - generally are not suitable for large-scale parallel processing. The fabrication of single-electron devices requires nanoscale geometrical arrangement of device components, that is, source and drain electrodes and Coulomb islands. Developing methods to fabricate nanoscale devices in large numbers at a time has been one of the major efforts of the nanotechnology community. A new study now demonstrates that this can be done with complete parallel processing using CMOS-compatible processes and materials. Furthermore, these single-electron devices can operate at room temperature, an essential requirement for practical implementations.
For nanoelectronics applications like single-electron devices to become practical, everyday items, they need to move from the highly individual and customized fabrication process typically found in laboratories to an automated, high-throughput and industrial-scale production environment. The reason this hasn't happened yet is that the various nanoscale pattern definition techniques used today - such as e-beam lithography, mechanically controllable break junctions, electromigration, electrodeposition, nanoscale oxidation, and scanning tunneling microscopy - generally are not suitable for large-scale parallel processing. The fabrication of single-electron devices requires nanoscale geometrical arrangement of device components, that is, source and drain electrodes and Coulomb islands. Developing methods to fabricate nanoscale devices in large numbers at a time has been one of the major efforts of the nanotechnology community. A new study now demonstrates that this can be done with complete parallel processing using CMOS-compatible processes and materials. Furthermore, these single-electron devices can operate at room temperature, an essential requirement for practical implementations.
Sep 18th, 2008
 Molecular-size motors have evolved in nature, where they are used in virtually every important biological process. In contrast, the development of synthetic nanomotors that mimic the function of these amazing natural systems and that could be used in man-made nanodevices is in its infancy. Building nanoscale motors is not just an exercise in scaling down the design of a macroworld engine to nanoscale dimensions. In addition to organic molecules, scientists increasingly are looking to DNA as a very promising way to fabricate nanomotors. The concept of a single DNA molecule nanomotor was already introduced in early 2002. However, this and subsequent designs require addition and removal of fuel and waste strands for motor function, although some artificial nanomotors can utilize alternative energy sources, including hydrolysis of the DNA backbone and ATP. Researchers at the University of Florida have now designed a photoswitchable single-molecule DNA nanomotor. It is the first fully reversible single-molecule DNA nanomachine driven by photons without any additional DNA strands as fuel.
Molecular-size motors have evolved in nature, where they are used in virtually every important biological process. In contrast, the development of synthetic nanomotors that mimic the function of these amazing natural systems and that could be used in man-made nanodevices is in its infancy. Building nanoscale motors is not just an exercise in scaling down the design of a macroworld engine to nanoscale dimensions. In addition to organic molecules, scientists increasingly are looking to DNA as a very promising way to fabricate nanomotors. The concept of a single DNA molecule nanomotor was already introduced in early 2002. However, this and subsequent designs require addition and removal of fuel and waste strands for motor function, although some artificial nanomotors can utilize alternative energy sources, including hydrolysis of the DNA backbone and ATP. Researchers at the University of Florida have now designed a photoswitchable single-molecule DNA nanomotor. It is the first fully reversible single-molecule DNA nanomachine driven by photons without any additional DNA strands as fuel.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





