Showing Spotlights 49 - 56 of 65 in category All (newest first):
 Studies have already shown the complexity of architecture that is achievable using DNA as building blocks. For instance, nanofabrication via molecular self-assembly has already resulted in simple DNA polyhedra with connectivities of a cube, octahedron, and a tetrahedron. Platonic solids - any one of five solids whose faces are congruent regular polygons and whose polyhedral angles are all congruent - are the most efficient at enclosing large volumes. The more complex the polyhedron, the greater its ability to encapsulate cargo, because the capsule size can be maximized while keeping the pore-size of the capsule minimal. The most complex platonic solid is the icosahedron and therefore this would be most suitable for achieving cargo encapsulation. The DNA shell of such a capsule could protect vulnerable drugs from degradation by proteins until they reach their target site. It could also prevent dangerous drugs from leaking out until the capsule reaches its intended target. The DNA shell could also allow attachment to a protein that could ferry the drug-loaded capsule to a target.
Studies have already shown the complexity of architecture that is achievable using DNA as building blocks. For instance, nanofabrication via molecular self-assembly has already resulted in simple DNA polyhedra with connectivities of a cube, octahedron, and a tetrahedron. Platonic solids - any one of five solids whose faces are congruent regular polygons and whose polyhedral angles are all congruent - are the most efficient at enclosing large volumes. The more complex the polyhedron, the greater its ability to encapsulate cargo, because the capsule size can be maximized while keeping the pore-size of the capsule minimal. The most complex platonic solid is the icosahedron and therefore this would be most suitable for achieving cargo encapsulation. The DNA shell of such a capsule could protect vulnerable drugs from degradation by proteins until they reach their target site. It could also prevent dangerous drugs from leaking out until the capsule reaches its intended target. The DNA shell could also allow attachment to a protein that could ferry the drug-loaded capsule to a target.
Apr 24th, 2009
 DNA nanomachines - synthetic DNA assemblies that switch between defined molecular shapes upon stimulation by external triggers - can be controlled by a variety of methods; these include pH changes and the addition of other molecular components, such as small molecule effectors, proteins and DNA strands. A team of Indian researchers has now taken structural DNA nanotechnology, where so far only in vitro applications have been demonstrated, across a new boundary and into living systems. They describe the successful operation of an artificially designed DNA nanomachine inside living cells, and show that these nanomachines work as efficiently inside cells as in vitro. The device is externally triggered by protons and functions as a pH sensor based on fluorescence resonance energy transfer (FRET) inside living cells.
DNA nanomachines - synthetic DNA assemblies that switch between defined molecular shapes upon stimulation by external triggers - can be controlled by a variety of methods; these include pH changes and the addition of other molecular components, such as small molecule effectors, proteins and DNA strands. A team of Indian researchers has now taken structural DNA nanotechnology, where so far only in vitro applications have been demonstrated, across a new boundary and into living systems. They describe the successful operation of an artificially designed DNA nanomachine inside living cells, and show that these nanomachines work as efficiently inside cells as in vitro. The device is externally triggered by protons and functions as a pH sensor based on fluorescence resonance energy transfer (FRET) inside living cells.
Apr 9th, 2009
 DNA, the fundamental building block of life, has become an intense nanotechnology research field. DNA molecules can serve as precisely controllable and programmable scaffolds for organizing functional nanomaterials in the design, fabrication, and characterization of nanoscale devices such as sensors and electronics. Most DNA research on controlled self-assembly deals with two-dimensional, i.e. flat, patterns and an expansion of these arrays into the third dimension has been challenging. New research coming out of UC Santa Barbara describes the self-assembly of multilayer hexagonal DNA arrays through highly regular interlayer packing. The researchers found that DNA arrays assembled into a two dimensional hexagonal pattern, or a sheet, assemble further into multilayer stacks.
DNA, the fundamental building block of life, has become an intense nanotechnology research field. DNA molecules can serve as precisely controllable and programmable scaffolds for organizing functional nanomaterials in the design, fabrication, and characterization of nanoscale devices such as sensors and electronics. Most DNA research on controlled self-assembly deals with two-dimensional, i.e. flat, patterns and an expansion of these arrays into the third dimension has been challenging. New research coming out of UC Santa Barbara describes the self-assembly of multilayer hexagonal DNA arrays through highly regular interlayer packing. The researchers found that DNA arrays assembled into a two dimensional hexagonal pattern, or a sheet, assemble further into multilayer stacks.
Jan 29th, 2009
 DNA, the fundamental building block of our genetic makeup, has become an intense nanotechnology research field. DNA molecules can serve as precisely controllable and programmable scaffolds for organizing functional nanomaterials in the design, fabrication, and characterization of nanometer scale electronic devices and sensors. The reason why DNA could be useful in nanotechnology for the design of electric circuits is the fact that it actually is the best nanowire in existence - it self-assembles, it self-replicates and it can adopt various states and conformations. The most basic and simplest form of DNA mechanical devices that are expected to be the first to demonstrate some close-to-reality functions are DNA tweezers. This concept was first introduced in 2000 by scientists at Bell Labs and Oxford University. To keep this type of tweezers running, two fuel DNA strands are alternately added to a buffered solution that contains the tweezers. These fuels are basically two stretches of complementary DNA, one of which closes the tweezers and the other opens them. The exciting potential applications for DNA tweezers include their use in constructing various molecular devices dedicated to repairing a functional unit in a cell, harnessing the delivery of drug molecules to pathogenic cells, or assembling nanoscale devices.
DNA, the fundamental building block of our genetic makeup, has become an intense nanotechnology research field. DNA molecules can serve as precisely controllable and programmable scaffolds for organizing functional nanomaterials in the design, fabrication, and characterization of nanometer scale electronic devices and sensors. The reason why DNA could be useful in nanotechnology for the design of electric circuits is the fact that it actually is the best nanowire in existence - it self-assembles, it self-replicates and it can adopt various states and conformations. The most basic and simplest form of DNA mechanical devices that are expected to be the first to demonstrate some close-to-reality functions are DNA tweezers. This concept was first introduced in 2000 by scientists at Bell Labs and Oxford University. To keep this type of tweezers running, two fuel DNA strands are alternately added to a buffered solution that contains the tweezers. These fuels are basically two stretches of complementary DNA, one of which closes the tweezers and the other opens them. The exciting potential applications for DNA tweezers include their use in constructing various molecular devices dedicated to repairing a functional unit in a cell, harnessing the delivery of drug molecules to pathogenic cells, or assembling nanoscale devices.
Nov 5th, 2008
 In case you haven't seen the absolutely amazing animation 'Cellular Visions: The Inner Life of a Cell' yet, go watch it now. In it, there is a sequence where a motor protein is sort of 'walking' along a filament, dragging this round sphere of lipids behind it. This kind of nanoscale biological motor is able to load/unload particular types of cargo without external stimuli, and transport them along cytoskeletal filaments by using the energy of adenosine triphosphate (ATP) hydrolysis within cells. Nanotechnology researchers are fascinated by the various molecular delivery systems that have evolved in nature and they are receiving increasing attention as blueprints for nanoscale actuators and building blocks to construct artificially-engineered bio-hybrid systems. Some researchers expect that artificial molecular transport systems which utilize microtubules motility will be an alternative way to pressure-driven or electrokinetic flow-based microfluidic devices. Researchers in Japan propose a molecular transport system that can achieve autonomous loading/unloading of specified cargoes. This system loads a cargo molecule through DNA hybridization.
In case you haven't seen the absolutely amazing animation 'Cellular Visions: The Inner Life of a Cell' yet, go watch it now. In it, there is a sequence where a motor protein is sort of 'walking' along a filament, dragging this round sphere of lipids behind it. This kind of nanoscale biological motor is able to load/unload particular types of cargo without external stimuli, and transport them along cytoskeletal filaments by using the energy of adenosine triphosphate (ATP) hydrolysis within cells. Nanotechnology researchers are fascinated by the various molecular delivery systems that have evolved in nature and they are receiving increasing attention as blueprints for nanoscale actuators and building blocks to construct artificially-engineered bio-hybrid systems. Some researchers expect that artificial molecular transport systems which utilize microtubules motility will be an alternative way to pressure-driven or electrokinetic flow-based microfluidic devices. Researchers in Japan propose a molecular transport system that can achieve autonomous loading/unloading of specified cargoes. This system loads a cargo molecule through DNA hybridization.
Apr 28th, 2008
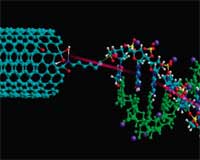 DNA, the blueprint of life, and electronics seem to be two completely different things but it appears that DNA could offer a solution to many of the hurdles that need to be overcome in further scaling down electronic circuits beyond a certain point. The reason why DNA could be useful in nanotechnology for the design of electric circuits is the fact that it actually is the best nanowire in existence - it self-assembles, it self-replicates and it can adopt various states and conformations. Not surprisingly, performing reliable experiments on a single oligo-DNA molecule is an extremely delicate task as partly contradicting research reports demonstrate: Different DNA transport experiments have shown that DNA may be insulating, semiconducting, or metallic. Among the numerous factors that could impact the results are the quality of the DNA-electrode interface, the base pair, the charge injection into the molecule, or environmental effects such as humidity or temperature. Researchers have now demonstrated a novel carbon nanotube-based nanoelectronic platform as proof of concept that single DNA molecules can be detected. This novel detection technique is based on change in electrical conductance upon selective hybridization of the complementary target DNA with the single stranded probe attached to the system. The single-stranded sequence-specific probe DNA whose ends are modified with amine is attached between two carbon nanotubes/nanowires using dielectrophoresis (DEP). This platform can be used for understanding how electrical charge moves through DNA which could help researchers understand and perhaps develop a technique for reversing the damage of DNA done by oxidation and mutation.
DNA, the blueprint of life, and electronics seem to be two completely different things but it appears that DNA could offer a solution to many of the hurdles that need to be overcome in further scaling down electronic circuits beyond a certain point. The reason why DNA could be useful in nanotechnology for the design of electric circuits is the fact that it actually is the best nanowire in existence - it self-assembles, it self-replicates and it can adopt various states and conformations. Not surprisingly, performing reliable experiments on a single oligo-DNA molecule is an extremely delicate task as partly contradicting research reports demonstrate: Different DNA transport experiments have shown that DNA may be insulating, semiconducting, or metallic. Among the numerous factors that could impact the results are the quality of the DNA-electrode interface, the base pair, the charge injection into the molecule, or environmental effects such as humidity or temperature. Researchers have now demonstrated a novel carbon nanotube-based nanoelectronic platform as proof of concept that single DNA molecules can be detected. This novel detection technique is based on change in electrical conductance upon selective hybridization of the complementary target DNA with the single stranded probe attached to the system. The single-stranded sequence-specific probe DNA whose ends are modified with amine is attached between two carbon nanotubes/nanowires using dielectrophoresis (DEP). This platform can be used for understanding how electrical charge moves through DNA which could help researchers understand and perhaps develop a technique for reversing the damage of DNA done by oxidation and mutation.
Dec 28th, 2007
 One of the many fascinating concepts in nanotechnology is the vision of molecular electronics. If realized, the shift in size from even the smallest computer chip today would be staggering - a quantum leap, so to speak (literally). Look at it this way: a single drop of water contains more molecules than the billions and billions of silicon chips ever produced. Molecular electronics engineers of tomorrow might use individual molecules to perform the functions in an electronic circuit that are performed by semiconductor devices today. Don't get your hopes up, though, that your next iPod will be truly nano. Scientists today are still struggling with the most basic requirements for molecular electronics, for instance, how to precisely and reliably position individual molecules on a surface. DNA-based nanostructuring is one approach that could lead to promising results. It has already been shown that DNA could be used to structure nanoscale surfaces. Now, a team in Germany has demonstrated that nanoscale objects of very different size can be deposited simultaneously and site-selectively onto DNA-displaying surfaces, based on sequence-specific DNA-DNA duplex formation.
One of the many fascinating concepts in nanotechnology is the vision of molecular electronics. If realized, the shift in size from even the smallest computer chip today would be staggering - a quantum leap, so to speak (literally). Look at it this way: a single drop of water contains more molecules than the billions and billions of silicon chips ever produced. Molecular electronics engineers of tomorrow might use individual molecules to perform the functions in an electronic circuit that are performed by semiconductor devices today. Don't get your hopes up, though, that your next iPod will be truly nano. Scientists today are still struggling with the most basic requirements for molecular electronics, for instance, how to precisely and reliably position individual molecules on a surface. DNA-based nanostructuring is one approach that could lead to promising results. It has already been shown that DNA could be used to structure nanoscale surfaces. Now, a team in Germany has demonstrated that nanoscale objects of very different size can be deposited simultaneously and site-selectively onto DNA-displaying surfaces, based on sequence-specific DNA-DNA duplex formation.
Aug 6th, 2007
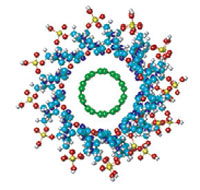 To achieve the full benefits of the amazing properties of carbon nanotubes (CNTs) researchers are exploring all kinds of CNT composite materials. Material engineers are interested because this will lead to lighter,stronger and tougher materials. Another fascinating area involves CNT/polymer composite structures that will lead to a vast range of improved and novel applications, from antistatic and EMI shielding to more efficient fuel and solar cells, to nanoelectronic devices. One particular area of CNT/polymer composites is dealing with DNA-CNTs hybrids. Although researchers expect a plethora of new applications, the fact that even the formation mechanism of these complexes is not yet clear shows how early in the game this research still is. This might be due to the fact that in spite of the quite large number of experimental investigations on the interaction between DNA and CNTs, the number of theoretical studies is limited. Researchers in Germany now present, for the first time, the results of a systematic quantum mechanical modeling of the stability and the electronic properties of complexes based on single-walled carbon nanotubes, which are helically wrapped by DNA molecules.
To achieve the full benefits of the amazing properties of carbon nanotubes (CNTs) researchers are exploring all kinds of CNT composite materials. Material engineers are interested because this will lead to lighter,stronger and tougher materials. Another fascinating area involves CNT/polymer composite structures that will lead to a vast range of improved and novel applications, from antistatic and EMI shielding to more efficient fuel and solar cells, to nanoelectronic devices. One particular area of CNT/polymer composites is dealing with DNA-CNTs hybrids. Although researchers expect a plethora of new applications, the fact that even the formation mechanism of these complexes is not yet clear shows how early in the game this research still is. This might be due to the fact that in spite of the quite large number of experimental investigations on the interaction between DNA and CNTs, the number of theoretical studies is limited. Researchers in Germany now present, for the first time, the results of a systematic quantum mechanical modeling of the stability and the electronic properties of complexes based on single-walled carbon nanotubes, which are helically wrapped by DNA molecules.
Jun 4th, 2007
 Studies have already shown the complexity of architecture that is achievable using DNA as building blocks. For instance, nanofabrication via molecular self-assembly has already resulted in simple DNA polyhedra with connectivities of a cube, octahedron, and a tetrahedron. Platonic solids - any one of five solids whose faces are congruent regular polygons and whose polyhedral angles are all congruent - are the most efficient at enclosing large volumes. The more complex the polyhedron, the greater its ability to encapsulate cargo, because the capsule size can be maximized while keeping the pore-size of the capsule minimal. The most complex platonic solid is the icosahedron and therefore this would be most suitable for achieving cargo encapsulation. The DNA shell of such a capsule could protect vulnerable drugs from degradation by proteins until they reach their target site. It could also prevent dangerous drugs from leaking out until the capsule reaches its intended target. The DNA shell could also allow attachment to a protein that could ferry the drug-loaded capsule to a target.
Studies have already shown the complexity of architecture that is achievable using DNA as building blocks. For instance, nanofabrication via molecular self-assembly has already resulted in simple DNA polyhedra with connectivities of a cube, octahedron, and a tetrahedron. Platonic solids - any one of five solids whose faces are congruent regular polygons and whose polyhedral angles are all congruent - are the most efficient at enclosing large volumes. The more complex the polyhedron, the greater its ability to encapsulate cargo, because the capsule size can be maximized while keeping the pore-size of the capsule minimal. The most complex platonic solid is the icosahedron and therefore this would be most suitable for achieving cargo encapsulation. The DNA shell of such a capsule could protect vulnerable drugs from degradation by proteins until they reach their target site. It could also prevent dangerous drugs from leaking out until the capsule reaches its intended target. The DNA shell could also allow attachment to a protein that could ferry the drug-loaded capsule to a target.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





