| Posted: Aug 18, 2014 |
Molecular learning machines under the microscope
|
|
(Nanowerk News) All human thoughts, feelings and actions are based on the fact that nerve cells communicate with each other. They pass impulses via synapses, intensify, weaken or block them. Like a ferry crossing a river, chemical messengers, the so-called neurotransmitters, pass from one side of the synapse to the other crossing a gap that is less than thousands of millimetres wide to dock to special synaptic receptors on the adjacent neuron. The molecular make-up of synaptic structures responsible for passing impulses is not fully known until today. Using a special imaging technique, scientists from the University of Würzburg have now succeeded in making processes at the nanoscale visible.
|
|
They present their results in the latest issue of Nature Communications ("Quantitative super-resolution imaging of Bruchpilot distinguishes active zone states").
|
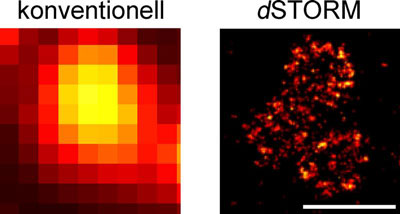 |
| Organisation of the Bruchpilot protein at active zones. High-resolution dSTORM imaging (right) shows details that cannot be visualised using conventional optical microscopy (left). Scale bars 500 nm. (Image: team Sauer / team Kittel)
|
|
Molecular machines in active zones
|
|
"The release of neurotransmitters in the so-called 'active zone' - a highly specialised sub-cellular region of the presynaptic neuron - is responsible for the information transfer at chemical synapses," explains Dr. Robert Kittel, who is the head of an Emmy Noether Group at the Department of Physiology of the University of Würzburg which focuses on the molecular mechanisms of synapses. Since 2009, he has studied the fruit fly Drosophila melanogaster to learn more about the active zone's physiology. He is particularly interested in the question of how structural changes of this active zone contribute to synaptic plasticity, thus enabling learning, for instance.
|
|
As Kittel puts it, "complex molecular machines" are at work in an active zone which convey the "extremely high spatial and temporal precisions of synaptic signal transduction". The neurotransmitter release is preceded by multi-stage processes in the active zone in the course of which the neutron provides the neurotransmitters in so-called vesicles. An exact image of the spatial distribution of the active zone's molecular constituents and thus information regarding the organisational principles has been the goal of Kittel's research for a long time, because it is these properties which are decisive for the function of the active zone. Collaborating with fellow Würzburg scientists, he has now moved closer to this goal.
|
|
Super-resolution optical microscopy
|
|
The breakthrough came with the cooperation with Professor Markus Sauer, head of the Department of Biotechnology and Biophysics at University of Würzburg's Biocenter. Together with his team, Sauer developed a method capable of delivering the desired images. Its name: dSTORM – direct Stochastic Optical Reconstruction Microscopy. A special form of high-resolution fluorescence microscopy, it enables imaging cellular structures and molecules with a ten- to hundred-fold increase in resolution compared to standard optical microscopy. The scale of the imaged objects is in the range of a few nanometres – that is, millionths of millimetres.
|
|
Electrophysiological measurements at the neurons of Drosophila combined with the dSTORM images then delivered the information the scientist had been looking for: the relationship between the spatial arrangement of special proteins at nanoscale and the functional properties of the active zone. In fact, this enabled the researchers to count the copies of the so-called "Bruchpilot" protein in the active zones and thereby work out quantitative structure-function relationships. "The analysis of the spatial organisation of molecules provides us information regarding the functional mechanisms of the active zone and helps us shed light on the basic mechanisms of brain function," is how the researchers interpret the result.
|

