| Posted: March 29, 2009 |
DNA-based assembly line for precision nano-cluster construction |
|
(Nanowerk News) Building on the idea of using DNA to link up nanoparticles scientists at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory have designed a molecular assembly line for predictable, high-precision nano-construction. Such reliable, reproducible nanofabrication is essential for exploiting the unique properties of nanoparticles in applications such as biological sensors and devices for converting sunlight to electricity. The work will be published online March 29, 2009, by Nature Materials.
|
|
The Brookhaven team has previously used DNA, the molecule that carries life’s genetic code, to link up nanoparticles in various arrangements, including 3-D nano-crystals. The idea is that nanoparticles coated with complementary strands of DNA — segments of genetic code sequence that bind only with one another like highly specific Velcro — help the nanoparticles find and stick to one another in highly specific ways. By varying the use of complementary DNA and strands that don’t match, scientists can exert precision control over the attractive and repulsive forces between the nanoparticles to achieve the desired construction. Note that the short DNA linker strands used in these studies were constructed artificially in the laboratory and don’t “code” for any proteins, as genes do.
|
|
|
|
Click on the image to download a high-resolution version.Using DNA to assemble nanoclusters: (a) (1) DNA linker strands (squiggly lines) are used to attach DNA-coated nanoparticles to a surface. (2) Linker strands are attached to the top side of the nanoparticle. (b) (3a) A nanoparticle of a second type with complementary DNA encoding recognizes the exposed linker strands and attaches to the surface-anchored nanoparticle. (4a and 5a) The assembled structure is released from the surface support, resulting in a two-particle, dimer cluster. (c) (3b) Alternatively, the immobilized particles produced in step (a) are released from the surface, leaving the opposite-side linker strands free to bind with multiple particles (4b) to form asymmetric "Janus" clusters.
|
|
The latest advance has been to use the DNA linkers to attach some of the DNA-coated nanoparticles to a solid surface to further constrain and control how the nanoparticles can link up. This yields even greater precision, and therefore a more predictable, reproducible high-throughput construction technique for building clusters from nanoparticles.
|
|
“When a particle is attached to a support surface, it cannot react with other molecules or particles in the same way as a free-floating particle,” explained Brookhaven physicist Oleg Gang, who led the research at the Lab’s Center for Functional Nanomaterials. This is because the support surface blocks about half of the particle’s reactive surface. Attaching a DNA linker or other particle that specifically interacts with the bound particle then allows for the rational assembly of desired particle clusters.
|
|
“By controlling the number of DNA linkers and their length, we can regulate interparticle distances and a cluster’s architecture,” said Gang. “Together with the high specificity of DNA interactions, this surface-anchored technique permits precise assembly of nano-objects into more complex structures.”
|
|
Instead of assembling millions and millions of nanoparticles into 3-D nanocrystals, as was done in the previous work, this technique allows the assembly of much smaller structures from individual particles. In the Nature Materials paper, the scientists describe the details for producing symmetrical, two-particle linkages, known as dimers, as well as small, asymmetrical clusters of particles — both with high yields and low levels of other, unwanted assemblies.
|
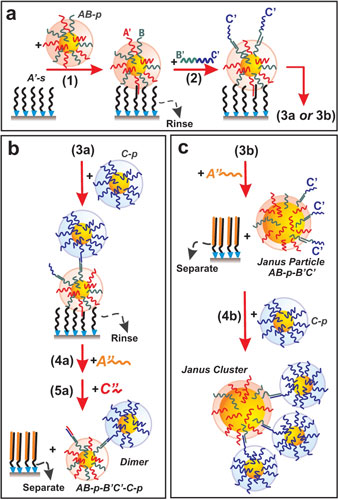 |
| Using DNA to assemble nanoclusters: (a) (1) DNA linker strands (squiggly lines) are used to attach DNA-coated nanoparticles to a surface. (2) Linker strands are attached to the top side of the nanoparticle. (b) (3a) A nanoparticle of a second type with complementary DNA encoding recognizes the exposed linker strands and attaches to the surface-anchored nanoparticle. (4a and 5a) The assembled structure is released from the surface support, resulting in a two-particle, dimer cluster. (c) (3b) Alternatively, the immobilized particles produced in step (a) are released from the surface, leaving the opposite-side linker strands free to bind with multiple particles (4b) to form asymmetric "Janus" clusters.
|
|
“When we arrange a few nanoparticles in a particular structure, new properties can emerge,” Gang emphasized. “Nanoparticles in this case are analogous to atoms, which, when connected in a molecule, often exhibit properties not found in the individual atoms. Our approach allows for rational and efficient assembly of nano-‘molecules.’ The properties of these new materials may be advantageous for many potential applications.”
nanoparticle dimers
|
|
For example, in the paper, the scientists describe an optical effect that occurs when nanoparticles are linked as dimer clusters. When an electromagnetic field interacts with the metallic particles, it induces a collective oscillation of the material’s conductive electrons. This phenomenon, known as a plasmon resonance, leads to strong absorption of light at a specific wavelength.
|
|
“The size and distance between the linked particles affect the plasmonic behavior,” said Gang. By adjusting these parameters, scientists might engineer clusters for absorbing a range of wavelengths in solar-energy conversion devices. Modulations in the plasmonic response could also be useful as a new means for transferring data, or as a signal for a new class of highly specific biosensors.
|
|
Asymmetric clusters, which were also assembled by the Brookhaven team, allow an even higher level of control, and therefore open new ways to design and engineer functional nanomaterials.
|
|
Because of its reliability and precision control, Brookhaven’s nano-assembly method would be scalable for the kind of high-throughput production that would be essential for commercial applications. Brookhaven Lab has applied for a patent on the assembly method as well as several specific applications of the technology. For information about the patent or licensing this technology, contact Kimberley Elcess at (631) 344-4151, or [email protected].
|
|
In addition to Gang, the team included materials scientist Dmytro Nykypanchuk, summer student Marine Cuisinier, and biologist Daniel (Niels) van der Lelie, all from Brookhaven, and former Brookhaven chemist Matthew Maye, now at Syracuse University. Their work was funded by DOE’s Office of Science and through a Goldhaber Distinguished Fellowship sponsored by Brookhaven Science Associates.
|
|
The Center for Functional Nanomaterials at BNL is one of the five DOE Nanoscale Science Research Centers (NSRCs), premier national user facilities for interdisciplinary research at the nanoscale. Together the NSRCs comprise a suite of complementary facilities that provide researchers with state-of-the-art capabilities to fabricate, process, characterize, and model nanoscale materials, and constitute the largest infrastructure investment of the National Nanotechnology Initiative. The NSRCs are located at DOE’s Brookhaven, Argonne, Lawrence Berkeley, Oak Ridge, and Sandia and Los Alamos National Laboratories.
|

