| Jul 08, 2013 |
Scientists make a distinctive step towards the understanding of bacterial resistance to ribosome-targeting antibiotics
|
|
(Nanowerk News) Researchers at the London Centre of Nanotechnology and Universities of Bristol and Southampton have made a distinctive step towards the understanding of bacterial resistance to ribosome-targeting antibiotics.
|
|
The ribosome – the supra-molecular machine responsible for the synthesis of protein in the cell – is the target of the majority of antibiotics used in medicine. In direct response bacteria have evolved a variety of strategies to overcome the action of antibiotics while maintaining the ability to synthesise proteins. One of these is to produce a protein named cfr – chloramphenicol/florfenicol resistance protein - that is able to subtly modify the ribosome structure to prevent the action of five different types of antibiotic and so make the bacteria resistant. Cfr was originally identified in bacteria from cattle, but has since spread to a variety of human pathogens including both methicillin-resistant Staphylococcus aureus (MRSA) and Escherichia coli.
|
|
Cfr is inherently interesting because its three-dimensional structure is not known and the modification that it produces involves a chemical reaction that is intrinsically unfavourable. To achieve this, Cfr reacts with two molecules of the same precursor to first “prime” itself with the components necessary for target modification and then to generate a free radical – a short-lived highly reactive compound – that is capable of modifying an unreactive target. The research team have investigated how Cfr uses the same catalytic machinery to drive these two different processes.
|
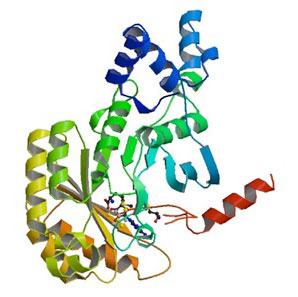 |
| X-ray structure of RImN – an enzyme closely related to Cfr - from Escherichia coli in complex with S-adenosyllmetionine (pdb entry: 3RFA).
|
|
Reported in PLoS ONE this week ("Cysteine Methylation Controls Radical Generation in the Cfr Radical AdoMet rRNA Methyltransferase"), the study shows that, although the “priming” process has no effect upon the ability of Cfr to recognise and bind the precursor, or upon the metal centre that is essential to Cfr activity, formation of the radical can occur only after priming has taken place. In this way Cfr controls the production of the reactive radical and prevents it from interfering with other functions within the bacterial cell.
|
|
Cfr binds tightly to the precursor molecule even when its ribosomal target is not present as oppose to other enzymes that belong to the same family. Thus it is possible that molecules with structures analogous to that of the precursor may disrupt Cfr activity and so restore antibiotic susceptibility in Cfr-producing bacteria.
|
|
A challenging outlook is to devise experimental methods that will enable such molecules to be easily identified and tested for their efficacy.
|

