| Apr 08, 2014 |
Synthetic gene circuits pump up cell signals
|
|
(Nanowerk News) Synthetic genetic circuitry created by researchers at Rice University is helping them see, for the first time, how to regulate cell mechanisms that degrade the misfolded proteins implicated in Parkinson’s, Huntington’s and other diseases.
|
|
The Rice lab of chemical and biomolecular engineer Laura Segatori has designed a sophisticated circuit that signals increases in the degradation of proteins by the cell’s ubiquitin proteasome system (UPS).
|
|
The research appears online today in Nature Communications ("Sensitive detection of proteasomal activation using the Deg-On mammalian synthetic gene circuit").
|
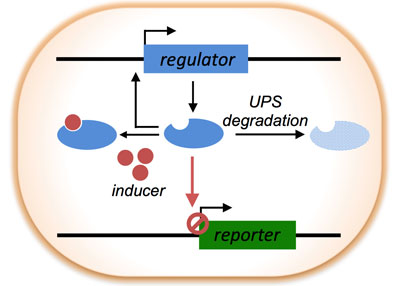 |
| The Deg-On system, an engineered regulator, controls the expression of a fluorescent reporter. Activation of the ubiquitin proteasome system (UPS) enhances degradation of the regulator and leads to an increase in fluorescent output. Tetracycline, an inducer, can be used to fine-tune the system and optimize detection of different levels of UPS activation. (Image: Segatori Group/Rice University)
|
|
The UPS is essential to a variety of fundamental cellular processes, including the cell cycle, DNA repair, immune response, cell death and the degradation of misfolded and damaged proteins. It has several components: ubiquitin molecules that tag misfolded proteins for degradation and proteasomes that latch onto the tagged proteins and break them down into harmless peptides.
|
|
When there are too few proteasomes in a cell, or they don’t function properly, misfolded proteins that remain floating in the cytoplasm can aggregate. These aggregates can form plaques, as often seen in the brains of people with neurodegenerative diseases.
|
|
The Rice team added to the cell a set of genetic circuits called Degradation On – “Deg-On” for short. These circuits produce a green fluorescent signal linked to UPS degradation and allow the researchers to monitor proteasomal activity.
|
|
“The overall goal is to develop a technology to screen for molecules that would enhance or activate degradation,” said Segatori, Rice’s T.N. Law Assistant Professor of Chemical and Biomolecular Engineering and an assistant professor of biochemistry and cell biology.
|
|
“The proteasome is essentially a big barrel that unfolds and chops up misfolded proteins. We know how to inhibit degradation, but we want to find ways to activate it, because we think that will be useful to help prevent accumulation of misfolded proteins and aggregation, which are associated with the development of a number of human diseases.”
|
|
The Deg-On circuit couples proteasomal degradation of an engineered tetracycline repressor to an easily detectable fluorescent signal. The tetracycline repressor is engineered to function as a UPS substrate; it essentially mimics a misfolded protein.
|
|
Normally, enhanced degradation would dampen the output signal, but this genetic circuit makes it possible to link enhanced degradation to an increase in output. The engineered repressor can still be regulated by the antibiotic tetracycline, which allows calibrating the system for the detection of even minimal activation of UPS degradation. An additional synthetic circuit provides a feedback loop that enables the self-amplification of the repressor so that adding tetracycline further boosts the signal.
|
|
|
|
The Rice team did extensive computer modeling of Deg-On to improve its sensitivity and dynamic range before building and testing the system on lab-standard HeLa cells. The team included graduate student and lead author Wenting Zhao, undergraduate Claire McWhite and Rice alumnus Matthew Bonem, in collaboration with Jonathan Silberg, an associate professor of biochemistry and cell biology at Rice.
|
|
“We came up with the idea of having a feedback loop, which is a self-activation loop for the tetracycline repressor, in our second circuit (an enhanced Deg-On),” Zhao said. “We found that a small increase in UPS activity caused a small decrease in the TetR repressor protein. Because TetR activates its own expression in the enhanced Deg-On, the fluorescent output signal is amplified and the circuit gains in sensitivity and dynamic range.”
|
|
“Wenting’s work was instrumental in predicting changes in the circuit architecture that would lead to enhanced sensitivity,” Segatori said.
|
|
The lab’s immediate goal is to create assays for the rapid detection of small molecules or genes that can increase proteasomal activity, she said. “This will help us rationally design compounds or strategies that could enhance degradation not only for the study and treatment of misfolding diseases but also for a variety of other applications.
|
|
“Misfolding and aggregation are among the main challenges in the fields of bioengineering and biotechnology. They are the bottleneck in the high-yield production of recombinant proteins, for example, in cells engineered to crank up expression of a protein of interest,” Segatori said.
|

