| Aug 03, 2014 |
Reprogramming bacteria as an intriguing approach to desalination
|
|
(Nanowerk News) Desalination has come a long way.
|
|
On Aug. 3, some 330 years ago, a certain Captain Gifford of His Majesty's Ship Mermaid, was asked to conduct onboard his 24-gun Royal Naval vessel what may have been the first government-sponsored, scientific desalination experiment.
|
|
Diarist and later Secretary to the Admiralty Commission in England Samuel Pepys wrote to Gifford saying, "Whereas a Proposal has been made to Us of an Engine to be fixed in one of Our Ships for the making an Experiment of producing fresh water (at Sea) out of Salt."
|
|
We do not know whether Gifford actually conducted the experiment, but we do know desalination--the pulling of salt, minerals and other contaminants from soil and water--has become a worldwide concern. Population increases, the scarcity of fresh water in arid regions and a greater need for environmental cleanup has scientists scrambling to improve the process.
|
|
Researchers at Rice University in Houston, Texas, for example, are computationally investigating ways to rewire one of desalination's most useful tools: Bacteria.
|
|
Bacteria as an environmental cleaning agent is based on the microorganisms' ability to sense its environment, consume pollutants, break them down and excrete different, less-harmful substances than the original contaminant. But bacteria's response mechanisms can do many other things such as provide scientifically discrete information, diagnose levels of toxins in food and water, detect poisonous chemicals, report dangerous compounds in the human body and more.
|
 |
| A view looking from the top down on a Bacillus subtillis colony in which conditions where such that an extracellular matrix--a mesh of proteins and sugars that can form outside bacterial cells--was produced, leading to biofilm formation. (Image: Hera Vlamakis, Harvard University Medical School)
|
|
That's why Jose Onuchic and Herbert Levine, co-directors of Rice's Center for Theoretical Biological Physics are working to treat bacteria like computers with the intention of reprograming them to perform specific activities.
|
|
The researchers have a plan to modify the proteins responsible for how bacteria respond to external stimuli, triggering the bacteria to predictably "decide" what actions to take when confronted with targeted environmental conditions.
|
|
Directed bacterial responses, the researchers believe, could revolutionize bacteria-based environmental cleanup, modern desalination and a host of medical and industrial applications.
|
|
The project, "Molecular Underpinnings of Bacterial Decision-Making" is one of a number of high-risk, potentially high-reward projects in the National Science Foundation's INSPIRE program. INSPIRE funds potentially transformative research that does not fit into a single scientific field, but crosses disciplinary boundaries.
|
|
"This research project by two highly respected scientists and their colleagues is an excellent example of basic research that can have tremendous societal benefits," says Kamal Shukla, program director in NSF's Division of Molecular and Cellular Biosciences.
|
|
The project is co-funded by NSF's Directorates for Biological Sciences and Mathematical and Physical Sciences.
|
|
Special molecules
|
|
"The information encoded in the genome not only contains the blueprint for making proteins that fold into unique 3-D structures," says Onuchic explaining the basis of the research, "but also contains rich information about functional protein-protein interactions." Two-component signaling (TCS) systems, found mainly in bacteria, are an example of this idea.
|
|
TCS systems are the dominant means by which bacteria sense the environment and carry out appropriate actions. These signaling pathways, determine how bacteria respond to heat, sunlight, toxins, oxygen and other environmental stimuli.
|
|
They also regulate characteristics such as how poisonous bacteria are, their ability to produce disease, their nutrient uptake, their ability to yield secondary organic compounds, etc.
|
|
"Our research tries to understand and potentially re-engineer two-component signaling systems," says Ryan Cheng, a postdoctoral fellow at Rice working on the project. "A successful understanding of the special molecules that make up these systems would allow us to take them apart like Lego blocks and start building new blocks or circuits to achieve a specific goal."
|
|
Earlier this year in a paper published in the Proceedings of the National Academy of Sciences ("Toward rationally redesigning bacterial two-component signaling systems using coevolutionary information"), the researchers revealed a scoring metric they devised to interpret how TCS proteins interact with each other and to predict how signaling modifications might affect TCS systems.
|
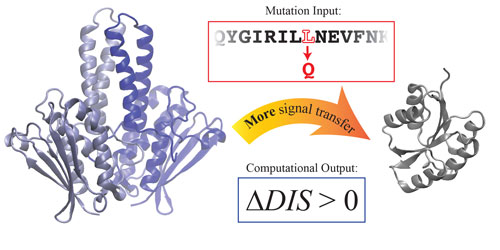 |
| Cheng et al. constructed a computational scoring function that can predict how single mutations can alter the interaction between two-component signaling (TCS) proteins. In this figure, positive scores correlate well with increased phosphotransfer ability between TCS pairs. (Image: Cheng et al., William Marsh Rice University)
|
|
The metric, based on sequence data from the coevolution of TCS proteins, could form a framework for fine tuning TCS signals and/or mix-matching TCS proteins leading to novel bacterial responses.
|
|
"Many proteins have evolved to produce specific behaviors under the additional constraint that they physically bind to another protein," says Faruck Morcos, a postdoctoral fellow at Rice, whose research focuses on computational biology and bioinformatics.
|
|
"Random mutations that may occur to one protein over geological timescales need to occur alongside mutations to the second protein in order to maintain their ability to interact with one another."
|
|
However, when the signal between two proteins that have evolved together is modified or a protein is matched with a non-evolutionary signaling partner, directed responses can occur.
|
|
"Hence, by applying methods from statistical physics, one can quantify and extract the statistical connections associated with amino acid coevolution between families of interacting proteins," Morcos says, and determine which proteins can successfully signal each other to produce predetermined outcomes.
|
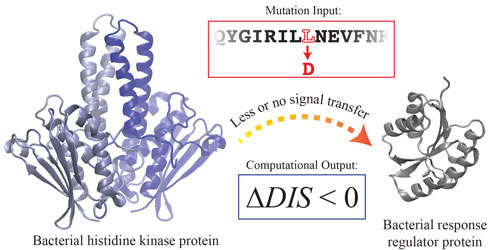 |
| Negative scores in the researchers scoring metric--depicted here--correlate with decreased phosphotransfer ability. The researchers say the scoring function alternately can be used to predict mutations that modify a wild type of interaction between two-component signaling proteins by a desired amount. (Image: Cheng et al., William Marsh Rice University)
|
|
Practical applications
|
|
With this operating premise, Onuchic and Levine, along with a small cadre of colleagues, plan to use the framework to engineer new, predictable behaviors in a model bacterium called Bacillus subtilis. Moreover, they plan to use B. subtilis as the prototype for changes in other protein-based systems.
|
|
"The potential applications for sanitation engineers are both numerous and profound," says Joshua Boltz, senior technologist and the biofilm technologies practice leader at CH2M HILL, a U.S. engineering company with major sewerage programs in London and Abu Dhabi, as well as clean water projects in the United States, Europe and Canada.
|
|
"Using membranes as a desalination tool to separate solids from liquids has emerged as a mature technology that is widely used globally," says Boltz zeroing in on an area where the research could benefit his industry. But, "A key concern with using membranes is their fouling, or a reduction in filtration capacity due to orifice clogging as a result of biofilms."
|
|
The researchers at Rice believe they can help reduce the buildup of biofilms in desalination equipment. Biofilms are thin layers of cells that stick to each other on a surface and have the ability to obstruct the flow of liquids in water purification systems.
|
|
"It has been shown experimentally that wrinkle formation in the biofilms of B. subtilis result from localized cell death," says Cheng. "Since cell death is regulated by two-component and related signaling systems, the potential for controlling the morphology and mechanical properties of biofilms exists."
|
|
The researchers surmise that this can perhaps be accomplished by introducing engineered bacteria to existing biofilms that can mechanically weaken existing biofilms through programmed cell death.
|
|
"While our research so far has exclusively dealt with quantifying the degree of interaction between a single pair of TCS proteins, a significant challenge will be to extend this work to make in vivo predictions," says Levine.
|
|
"Extending our methodology to complicated systems containing many potentially competing protein-protein interactions, e.g. living systems, will be a significant challenge for us in the future. We hope to extend this methodology to predictively understand how making a specific site-directed mutation affects the characteristics of an organism."
|



