| Jun 02, 2015 |
Intelligent bacteria for detecting disease
|
|
(Nanowerk News) Another step forward has just been taken in the area of synthetic biology. Research teams from Inserm and CNRS (French National Centre for Scientific Research) Montpellier, in association with Montpellier Regional University Hospital and Stanford University, have transformed bacteria into "secret agents" that can give warning of a disease based solely on the presence of characteristic molecules in the urine or blood. To perform this feat, the researchers inserted the equivalent of a computer programme into the DNA of the bacterial cells. The bacteria thus programmed detect the abnormal presence of glucose in the urine of diabetic patients.
|
|
This work, published in the journal Science Translational Medicine ("Detection of pathological biomarkers in human clinical samples via amplifying genetic switches and logic gates"), is the first step in the use of programmable cells for medical diagnosis.
|
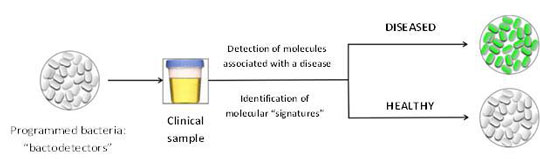 |
| Principle of the use of modified bacteria for medical diagnosis.
|
|
Bacteria have a bad reputation, and are often considered to be our enemies, causing many diseases such as tuberculosis or cholera. However, they can also be allies, as witnessed by the growing numbers of research studies on our bacterial flora, or microbiota, which plays a key role in the working of the body. Since the advent of biotechnology, researchers have modified bacteria to produce therapeutic drugs or antibiotics. In this novel study, they have actually become a diagnostic tool.
|
|
Medical diagnosis is a major challenge for the early detection and subsequent monitoring of diseases. "In vitro" diagnosis is based on the presence in physiological fluids (blood and urine, for example) of molecules characteristic for a particular disease. Because of its noninvasiveness and ease of use, in vitro diagnosis is of great interest. However, in vitro tests are sometimes complex, and require sophisticated technologies that are often available only in hospitals.
|
|
This is where biological systems come into play. Living cells are real nano-machines that can detect and process many signals and respond to them. They are therefore obvious candidates for the development of powerful new diagnostic tests. However, they have to be provided with the appropriate "programme" for them to successfully accomplish the required tasks.
|
|
To do this, Jérôme Bonnet's team in Montpellier's Centre for Structural Biochemistry (CBS) had the idea of using concepts from synthetic biology derived from electronics to construct genetic systems making it possible to "programme" living cells like a computer.
|
|
The transcriptor: the cornerstone of genetic programming
|
|
The transistor is the central component of modern electronic systems. It acts both as a switch and as a signal amplifier. In informatics, by combining several transistors, it is possible to construct "logic gates," i.e. systems that respond to different signal combinations according to a predetermined logic. For example, a dual input "AND" logic gate will produce a signal only if two input signals are present. All calculations completed by the electronic instruments we use every day, such as smartphones, rely on the use of transistors and logic gates.
|
|
During his postdoctoral fellowship at Stanford University in the United States, Jérôme Bonnet invented a genetic transistor, the transcriptor.
|
|
The insertion of one or more transcriptors into bacteria transforms them into microscopic calculators. The electrical signals used in electronics are replaced by molecular signals that control gene expression. It is thus now possible to implant simple genetic "programmes" into living cells in response to different combinations of molecules .
|
|
In this new work, the teams led by Jérôme Bonnet (CBS, Inserm U1054, CNRS UMR5048, Montpellier University), Franck Molina (SysDiag, CNRS FRE 3690), in association with Professor Eric Renard (Montpellier Regional University Hospital) and Drew Endy (Stanford University), applied this new technology to the detection of disease signals in clinical samples.
|
|
Clinical samples are complex environments, in which it is difficult to detect signals. The authors used the transcriptor's amplification abilities to detect disease markers, even if present in very small amounts. They also succeeded in storing the results of the test in the bacterial DNA for several months.
|
|
The cells thus acquire the ability to perform different functions based on the presence of several markers, opening the way to more accurate diagnostic tests that rely on detection of molecular "signatures" using different markers.
|
|
"We have standardised our method, and confirmed the robustness of our synthetic bacterial systems in clinical samples. We have also developed a rapid technique for connecting the transcriptor to new detection systems. All this should make it easier to reuse our system," says Alexis Courbet, a postgraduate student and first author of the article.
|
|
As a proof of concept, the authors connected the genetic transistor to a bacterial system that responds to glucose, and detected the abnormal presence of glucose in the urine of diabetic patients.
|
|
"We have deposited the genetic components used in this work in the public domain to allow their unrestricted reuse by other public or private researchers, " says Jérôme Bonnet.
|
|
"Our work is presently focused on the engineering of artificial genetic systems that can be modified on demand to detect different molecular disease markers," he adds. In future, this work might also be applied to engineering the microbial flora in order to treat various diseases, especially intestinal diseases.
|

