| Dec 21, 2012 |
How molecular transports change gear
|
|
(Nanowerk News) The motor protein myosin-V, which hauls molecular cargoes around cells by ratcheting along filaments of actin, switches between two different molecular mechanisms of movement depending on the environment. This finding by a research group led by Toshio Yanagida of the RIKEN Quantitative Biology Center, Osaka, and Osaka University, could form the basis for designing energy-saving artificial nano-motors ("Switching of myosin-V motion between the lever-arm swing and Brownian search-and-catch").
|
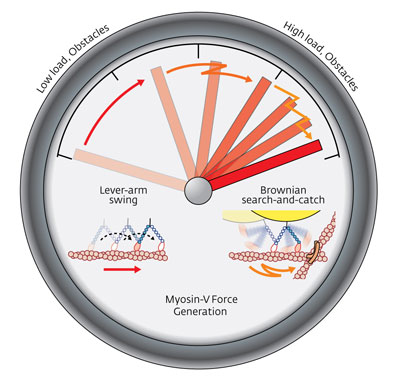 |
| As the molecular load on myosin-V increases, lever-arm swing motion (left) switches to Brownian search-and-catch (right).
|
|
Previous work by other researchers showed that myosin-V typically uses the two mechanisms—lever-arm swing and Brownian search-and-catch—alternately to propel itself along a filament hand-over-hand. Myosin-V possesses two arm-like projections, the heads of which bind to actin. When myosin-V links to the energy storage molecule adenosine triphosphate (ATP) the rear head detaches from the actin filament. As ATP releases energy by losing a phosphate, the front head then goes through a lever-arm swing motion pulling forward on the filament like an oar through water, dragging the rear projection with it. The rear head then swings over and forward while buffeted by passing molecules in random Brownian motion. As it nears the filament in front, it catches onto it.
|
|
Yanagida and his colleagues, including Keisuke Fujita and Mitsuhiro Iwaki, were able to attach a fluorescent polystyrene bead to the rear projection of myosin-V with a strand of DNA. This allowed them to measure the motions of the motor molecule accurately by tracking the displacement of the bead. They could also measure the force each head exerted by trapping the bead and holding it steady using the laser light mechanism known as optical tweezers under different loads and environmental conditions.
|
|
The results showed that for low loads along filaments where there are no obstacles, the bulk of the work of myosin-V’s motion is executed by the lever-arm swing mechanism. But at higher loads, and in less predictable environments, the force capable of being exerted by lever-arm swing reaches a maximum and Brownian search-and-catch motion automatically takes over (Fig. 1). Cells contain a meshwork of crisscrossing actin filaments and there is always the possibility of colliding with moving molecules and vesicles to hinder the transport of myosin-V’s molecular cargoes. Under these circumstances the ‘high-stepping’ Brownian search-and-catch motion comes into its own.
|
|
“We are hopeful that the studies of other biological actuators or simulations will show that our theory for myosin-V movement is universal and therefore adds a much more concrete paradigm to the design of artificial nano-machines,” says Yanagida.
|

