| Posted: Nov 08, 2006 | |
Molecular engineering: networks of nanotubes and containers |
|
| (Nanowerk Spotlight) Back in 2001, Swedish researchers developed techniques for creating complex two- and three-dimensional networks of nanotubes and µm-sized containers from liquid crystalline lipid bilayer materials based on the propensity in liposomes to undergo complex shape-transitions under mechanical excitations. The membrane composition and container contents can be controlled allowing chemical programming of networks in studies of enzyme kinetics, reaction-diffusion phenomena, and single-biomolecule detection. Materials contained in the networks can be routed among containers. Thus, networks of nanotubes and vesicles serve as a platform to build nanofluidic devices operating with single molecules and particles and offer new opportunities to study chemistry in confined biomimetic compartments. The networks can furthermore be used to build nanoscale chemical laboratories for applications in analytical devices as well as to construct computational and complex sensor systems that can also be integrated to living cells. In recent work, the researchers have now demonstrated that these nanotube-container networks can be constructed directly from plasma membranes of cultured cells. | |
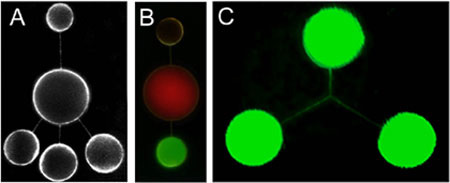 |
Networks of nanotubes and containers formed using mechanical excitation. The nanotubes are between 100 and 200 nm in radius and the containers 4-10 µm in diameter. A shows an open, five-liposome network stained with a fluorescent membrane dye. In B, the contents of the three nanotube-connected liposomes have been differentiated using fluorescent dyes and fluorescent colloidal particles. C shows a three-way nanotube junction. (Image: Orwar Research Group) |
| Plasma membrane vesicles in itself are not a new concept, and people have done different studies on them regarding composition and functionality, mainly by isolating pinched-off vesicles in a large batch. The new aspect of this recent work done at the Orwar Research Lab is to be able to visualize membrane proteins on one or more single vesicles, and being able to create different network structures by micromanipulation, similar to the liposome networks developed previously. | |
| Nanotube conjugated cells also exist in nature, and evidence has been found that cells can exchange materials through such channels. In the laboratory, though, an important challenge in constructing nanotube-vesicle networks has been to obtain membrane proteins and lipids from natural sources in high yields and with maintained functionality. The most elegant approach would be to form the nanotube-vesicle networks directly from a native cell membrane; which is exactly what the Swedish scientists did. | |
| Specifically, they used the fact that, under certain conditions, cells can form unilamellar protrusions, also known as membrane blebs. Such structures can serve as ideal precursors for nanotube-vesicle network formation as they are compatible with the micromanipulation tools used for synthetic vesicles. | |
| Applying a combination of dithiothreitol (DTT) and formaldehyde, the formation of micronsized membrane blebs is induced. Using the microelectroinjection technique developed earlier the researchers demonstrate that membrane blebs can be used to form surface adhered networks of plasma membrane vesicles. | |
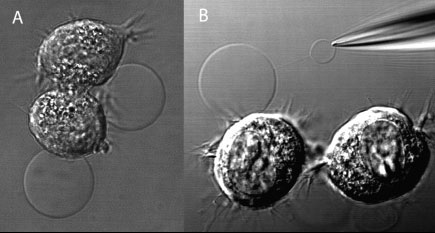 |
(A) DIC image of adherent NG108-15 cells, displaying membrane blebs induced by a combination of DTT and formaldehyde. (B) Subsequently, one bleb was electroinjected with a buffer-filled micropipet. Following translation of the micropipet, a nanotube connection is formed. Buffer injection leads to the formation of a daughter vesicle at the micropipet tip. (Reprinted with permission from the American Chemical Society) |
| This method provides high yields, and composition, orientation, as well as function are conserved. Furthermore, given that only about 250 µm2 of membrane is needed to build a three vesicle network (given a 5 µm vesicle diameter and 40 µm total tube length), such a network can be built from a single cell. | |
| "The fact that we can now create these networks with cellular plasma membrane components is exciting and opens up a new range of possibilities of probing cellular processes in these simplified structures" Brigitte Bauer explains to Nanowerk. "We can functionalize nanotube vesicles with desired proteins and lipids, where it is possible to study transport activity across nanotubes and membranes, as well as to study substrate conversion down to the single-molecule limit." | |
| Bauer, a student in the Orwar Research Group at the Chemical and Biological Engineering department at Chalmers University of Technology in Sweden, is first author of a recent paper, titled "Direct Reconstitution of Plasma Membrane Lipids and Proteins in Nanotube-Vesicle Networks", published online on the September 29, 2006 issue of Langmuir. | |
| "Now, that we can create nanotube-vesicle networks that originate from the plasma membrane of single cells, we are able to probe membrane proteins on a single molecule level" says Bauer. "The cells can be cherry-picked from a tissue or a culture to build a nanotube-vesicle network with a desired membrane makeup." | |
| Bauer points out that it is of great interest to develop biomimetic systems that are simple-to-use models to study events on a small scale level, as they happen in a single cell. Being able to use single cells as a material source greatly extends the application range for these systems, allowing the study of complex cellular processes in a simplified environment. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
