| Posted: Jun 11, 2009 | |
Nanoscale piston |
|
| (Nanowerk Spotlight) Reciprocating devices are a common part of the macroscopic world. Examples of reciprocating machines are petrol and diesel engines or a hydraulic pump. At the core of these machines is a piston and cylinder assembly where the piston executes a reciprocating motion inside the cylinder. | |
| Reciprocating motion like that in a piston has not been available in a nanoscale machine until now. Ned Seeman and his team at New York University have designed a DNA device that exhibits reciprocal motion. | |
| "We have used the PX-JX2 device, a robust sequence-dependent nanomechanical DNA machine, as the basis for constructing a pair of reciprocal devices, wherein one device assumes one state, while the other device assumes the opposite state," Seeman explains to Nanowerk. "This capability has not been demonstrated yet on the nanomolecular scale. The key point of these two PX-JX2 devices is that they share the same set strands, so that they make reciprocal motions synchronously. With the same control strands, one device is set to the PX state, and the other is set to the JX2 state; the opposite set strands switch the two states." | |
 |
|
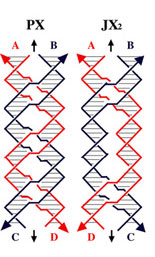
| PX and JX2 motifs of DNA | |
| For more on the PX-JX2 device see this section on Seeman's website or read about a specific example we covered in a previous Nanowerk Spotlight ("Nanorobotic arm to operate within DNA sequence"). | |
| Seeman says that the strategy of using the same control strands to set distinct states of two devices lends a new element to the toolbox for nanorobotics and for machines whose states are set by computational processes. | |
| However, this novel device is just a demonstration in principle. Just don't expect DNA machines with thousands of rpm yet. The synchronous reciprocal motion demonstrated by Seeman's team, i.e. a complete cycle of operation to reach equilibrium, took almost half a day. | |
| Another key remaining issue is putting these devices into a programmed array. | |
| The researchers have published their findings in the June 4, 2009 online edition of Nano Letters ("Reciprocal DNA Nanomechanical Devices Controlled by the Same Set Strands"). | |
| The formation, transformation, and reciprocal motion of the devices have been demonstrated by nondenaturing gel electrophoresis as well as direct observation through atomic force microscopy (AFM). | |
| "To obtain direct visualization of the transformation and of the reciprocal motion of the constructed devices, we have designed a one-dimensional system visible by AFM," Seeman explains. "We connect a series of DNA PX triangles with the devices in the PX or JX2 states using a pair of doubly cohesive sticky-ends. The PX triangle, containing 76 nucleotide pairs per edge, is large enough to be observed by AFM and allowed us to observe the reciprocal motion of the device." | |
| Seeman points out that the strategy of using the same control strands to set distinct states of two devices enables many nanoscale capabilities from reciprocating machines to molecular choreography. | |
| "The ability to correlate the motions of molecular devices can lead to complex behavior on the nanoscale" he says. "For example, in a recent device the legs of a bipedal walker (see "A Bipedal DNA Brownian Motor with Coordinated Legs") communicate with each other, producing an autonomous walker." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
