| Posted: May 07, 2010 | |
Re-shaping bacterial inclusion bodies as improved nanomaterials for tissue engineering |
|
| (Nanowerk Spotlight) Cultured mammalian cells prefer growing on structured rather than on completely flat surfaces. In regenerative medicine, in which human cells must grow on artificial scaffolds to replace damaged tissue, appropriate biological signals, such as growth factors, but also mechanic stimuli should be provided at the nano and microscales for cell attachment and proliferation, mimicking the natural cell matrices in organic tissues. The straightforward fabrication of nanostructured surfaces as scaffolds for tissue engineering is complex, but instead, micro- and nanorugosities can be easily generated on flat surfaces by either top-down or bottom-up approaches. | |
| Regarding top-down procedures, lithographic methods are extremely convenient for the generation of nano environments with highly regular patterns (mainly grooves or pits, with defined deeps and 2D distribution). Surfaces modified in such a way have been successfully exploited in the evaluation of cell responses in front of different nano environments, from morphological, architectonic and transcriptomic points of view ("The control of human mesenchymal cell differentiation using nanoscale symmetry and disorder"). | |
| The top-down approximation to nano environment engineering is, however, strongly limited by the nature of the used material, a fact that restricts the in vivo applicability of this approach. For medical applications, bottom-up strategies seems to be more feasible, as essentially any scaffold material can be decorated with micro- and nanoparticles of appropriate sizes and at optimal densities. Ceramic, polymeric and metal particles have so far explored for this purpose, although with rather irregular results. | |
 |
|
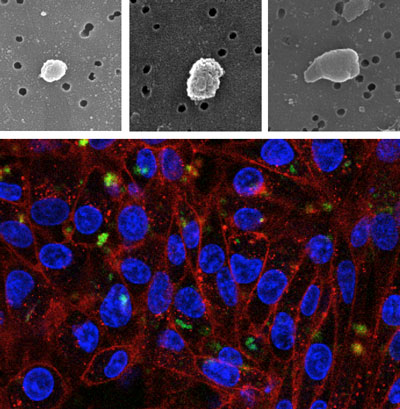
| Top: Scanning electron microcopy of inclusion bodies purified from E. coli wild type (left), DnaK- (center) and ClpP- (right). Bottom: HeLa cells growing on polystyrene surfaces decorated with inclusion bodies formed by a fluorescent GFP variant. Green signals: fluorescent inclusion bodies; blue signals: cell nucleus; red signals: cell membranes. (Images: Elena García-Fruitós, Joaquín Seras-Franzoso, Esther Vazquez) | |
| Recently, we determined that bacterial inclusion bodies, pseudo-spherical protein aggregates often observed in recombinant bacteria, can be used as micro- or nanoparticulate materials for surface decoration and the stimulation of cell proliferation. | |
| Inclusion bodies emerge in the bacterial cytoplasm as a result of the overproduction of foreign polypeptides in microbial cell factories, that being unable to reach their native conformation tend to aggregate as insoluble protein clusters. These particles have been traditionally observed as obstacles in protein production processes, which are obviously aimed to obtain soluble products usable in industry and therapy. | |
| Ranging in size between 50 and 500nm, inclusion bodies are mechanically stable, nanoparticulate entities that because of their proteinaceous composition, are fully biocompatible. | |
| In cooperation with Prof. Jaume Veciana from the Institute of Material Sciences of Barcelona (ICMAB-CSIC), we have demonstrated that bacterial inclusion bodies formed by biologically irrelevant polypeptides (such as GFP), are convenient biomaterials for the bottom-up decoration of substrates for mammalian cell growth, dramatically favoring cell attachment and growth through the mechanical stimuli exerted on mammalian cells ("Surface Cell Growth Engineering Assisted by a Novel Bacterial Nanomaterial"). | |
| Inclusion bodies can be easily produced and purified from recombinant Escherichia coli by cost-effective procedures, and straightforward used as biomaterials upon simple washing steps. | |
| In a step further, we have explored the genetic background of the producing bacterial cells in order to identify specific genes responsible for the biological fabrication of inclusion bodies, in an attempt to tailor properties of these particles relevant to their role as topographical modifiers. | |
| In this regard, it must be noted that bacterial inclusion bodies are generated in the context of the quality control network, a complex cell system formed by chaperones and proteases with synergistic, antagonistic and overlapping activities. Recombinant proteins that are reluctant to folding attempts mediated by chaperones either aggregate or are degraded by proteases in chaperone-driven processes. Interestingly, polypeptides deposited in inclusion bodies are continuously removed from these particles by disaggregating chaperones (DnaK, DnaJ, ClpB, IbpA, IbpB) working coordinately with ClpP and Lon proteases. | |
| In our study, we explored the formation of inclusion bodies in several bacterial strains lacking key chaperones or proteases, observing abnormal shapes in inclusion bodies formed in a ClpP-deficient E. coli background. The absence of this chaperone results in tear-shaped protein particles of around 400 nm in diameter, that when used to decorate polystyrene surfaces for mammalian cell culture resulted in enhanced cell growth stimulation properties, specially observed in PC12 neuronal cells ("Tunable geometry of bacterial inclusion bodies as substrate materials for tissue engineering" – free article). | |
| The unexpected possibility to genetically define the geometry of inclusion bodies – usually spherical – as particulate entities during their in vivo fabrication process, and the consequent echo of these modifications on biological interfaces open intriguing possibilities for the further development and improvement of these novel biomaterials in tissue engineering or other potential applications. | |
| By Elena García-Fruitós, Joaquin Seras-Franzoso, Esther Vazquez and Antonio Villaverde. CIBER en Bioingeniería, Biomateriales y Nanomedicina and Institut de Biotecnologia i de Biomedicina and Departament de Genética i de Microbiologia, Universitat Autónoma de Barcelona, 08193 Bellaterra, Barcelona, Spain | |
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
