| Posted: Jun 02, 2008 | |
Integrating biological functionality into plastic surfaces |
|
| (Nanowerk Spotlight) The controlled patterning of surfaces with biomolecules is of great importance for future generations of micro and nano biodevices (e.g. biochips, BioMEMS, lab-on-a-chip) and biomaterials. Even with current state-of-the-art technology, this patterning requirement, i.e. the immobilization and controlled and precise placement of biomolecules, often is a limiting step in the fabrication process. | |
| Commonly applied substrate materials for such biodevice applications are inexpensive polymers; but polymer surfaces are complex to chemically pattern in larger numbers. By combining two known techniques, micro-contact printing and injection molding in a new, innovative way, researchers in Denmark have now demonstrated a surprisingly successful methodology for transferring micro- and nanoscopic patterns of functionally active proteins to polymer surfaces during injection molding of hot polymer melt. | |
| "Previous work has documented the challenges in making biomolecule patterns with feature sizes of less than 350 nm using inexpensive scalable soft lithography techniques such as direct microcontact printing" Dr. Niels Bent Larsen explains to Nanowerk. "Our novel in-mold patterning technique easily produced patterns spanning three orders of length scales without any indication that the 310 nm size of the smallest features presented is a lower limit to the feature size. The broad span of usable polymers and feature sizes allows for applications in areas ranging from cell culture to nanobiosensors." | |
| Larsen, a research professor in the Department of Micro- and Nanotechnology at the Technical University of Denmark (DTU), together with his collaborators from DTU demonstrated that biotinylated biomolecules can be patterned on surfaces from solution after in-mold patterning of streptavidin, employing the widely used streptavidin-biotin system. | |
| The scientists published their findings in the April 17, 2008 online edition of Advanced Materials ("Protein In-Mold Patterning"). | |
| Larsen explains that his team's work has resulted in two core findings: | |
| First, a scientific finding that a range of proteins typically denaturating above 60 °C may survive exposure to very high temperatures, around 250 °C, in a functionally active form if the exposure time is extremely short, i.e. nanoseconds. | |
| Ultra-fast folding and unfolding of proteins has been explored thoroughly during the past decade, by the group of Alan R. Fersht in Cambridge,UK ("The complete folding pathway of a protein from nanoseconds to microseconds") and others, mostly in solution or by computer simulation. Recently, denaturation of selected proteins by ultra-short heat pulses (< 50 nanoseconds) up to 230 °C has also been probed using specialized pulsed laser equipment in the group of Cristobal G. dos Remedios at University of Sydney ("Nanosecond Responses of Proteins to Ultra-High Temperature Pulses "). | |
| "Our work adds to this existing pool of knowledge by demonstrating, for the first time to our knowledge, that a range of commonly used proteins in bioanalysis and cell culture can remain functionally active at even higher temperatures and in a very useful process for engineering nanotechnology, rather than in the lab on an optical breadboard." | |
| The second, scientific and technological, core finding of Larsen's team is that a single monolayer of proteins adsorbed to a metal surface can be transferred with high efficiency into the surface layer of solidifying polymer, remain there, and still be biologically active. | |
| "Many things could go wrong in that process, including that the protein would prefer to remain on the metal surface, that the protein would get buried in the polymer melt, or that it would be only be adhering weakly to the polymer matrix and therefore be removed during later washing steps of bioanalysis or cell culture" says Larsen. "Apparently we hit a lucky compromise of nature where all the properties we wished for were fulfilled." | |
 |
|
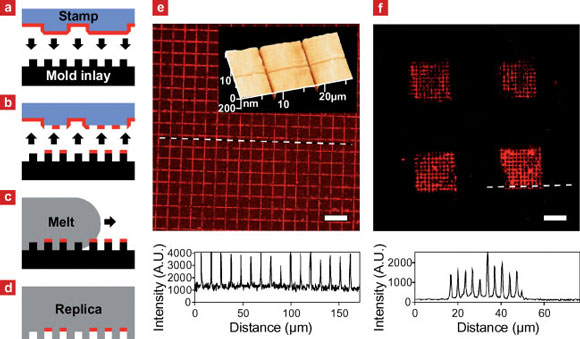
| a?d) Outline of the in-mold patterning process. a) Biomolecules adsorbed from solution are transferred from the raised areas of an elastomer stamp onto b) elevated regions of a mold inlay by microcontact printing. c) Injection of hot polymer melt over the surface of the cold mold inlay results in d) the simultaneous shaping of the replica surface and immobilization of the biomolecule pattern in recessed areas of the replica. e) Fluorescence microscopy of fluorescently labeled IgG transferred onto a PMMA replica by injection molding. The intensity profile reveals a grid line spacing equal to that of the mold inlay surface relief. Insert: atomic force microscopy of the replica surface shows an equivalent grid of trenches consistent with the transfer of IgG to recessed surface areas. f) Fluorescence microscopy image and line intensity profile at the dashed line of a polystyrene replica patterned with fibronectin (FN) at multiple length scales: A stamp with 40 x 40 µm2 raised surface areas, laterally separated by 40µm, transferred FN to a mold inlay with 310nm wide lines separated by 3100 nm. After transfer of FN by injection molding the replica was immunolabeled using primary and fluorescently labeled secondary antibodies. Scale bars: 20µm. (Reprinted with permission from Wiley-VCH Verlag) | |
| The Danish team employed microcontact printing using flat or structured stamps for the transfer of proteins to the mold inlay surface. Larsen describes the process, illustrated in above schematic: The targeted protein was adsorbed onto a poly(dimethyl siloxane) (PDMS) stamp from solution. The protein-coated stamp was brought into conformal contact with the raised areas of a topographically structured mold inlay and removed again, leaving proteins where the stamp and inlay were in contact. Hot polymer melt was injected and after cooling times from seconds up to one minute, the solidified shaped replica with the protein pattern immobilized at recessed surface areas was ejected from the mold. | |
| "Going into this field originated with our long-time search for new ways of controlling cell behavior in human cell cultures by surface nano-shape (nano-topography) or surface nano-patterns of biomolecules" Larsen explains the motivation for the team's work. "We pursued both topography and chemistry in parallel, nano-topography by injection molding and nano-chemistry by microcontact printing. Thermal modeling of the injection molding process at nanometer length scales revealed the extremely fast cooling rates at the metal mold surface, and this lead to the idea of combining injection molding and microcontact printing of proteins." | |
| There appears to be a range of potential applications for this work in the fields of cell culture and bioanalysis. Conventional cell culture relies heavily on the addition of blood serum extracts to sustain cell viability. However, blood serum is a mix of many known and unknown components so there is a large impetus for switching to serum-free cell culture. | |
| Larsen points out that the ability to built-in selected cell-supporting proteins in the surface of disposable cell culture plastics in a reproducible and inexpensive process has great commercial appeal. "Bioanalytical applications could also benefit greatly from built-in homogeneous or nanopatterned anchoring areas for particular sensing molecules" he says. "A widely used system is the anchoring of biotin-marked sensing molecules to surface immobilized streptavidin molecules, which we demonstrate to work smoothly after In-Mold Patterning of the streptavidin." | |
| The concepts outlined above are currently being commercialized by a Danish start-up company, InMold Biosystems. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
