Showing Spotlights 313 - 320 of 544 in category All (newest first):
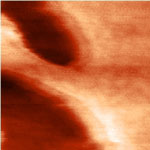 Lipids are the main component of the outermost membrane of cells. Their role is to seperate the inner and outer media of the cell and prevent any ionic current between these two media. Because of this last property, lipid layers can be thought of as good ultra-thin insulators that could be used in the development of electronic devices. So far though, because of their inherent instability in air, their use in advanced processes has been limited. This might change, though, since researchers in France have shown the possibility to stabilize by polymerization a lipid monolayer with a thickness of 2.7 nm directly at the surface of H-terminated silicon surface therefore opening a whole new world of possibilities of the use of these layers. Now, they reported the electrical performance of stabilized lipid monolayers on H-terminated silicon.
Lipids are the main component of the outermost membrane of cells. Their role is to seperate the inner and outer media of the cell and prevent any ionic current between these two media. Because of this last property, lipid layers can be thought of as good ultra-thin insulators that could be used in the development of electronic devices. So far though, because of their inherent instability in air, their use in advanced processes has been limited. This might change, though, since researchers in France have shown the possibility to stabilize by polymerization a lipid monolayer with a thickness of 2.7 nm directly at the surface of H-terminated silicon surface therefore opening a whole new world of possibilities of the use of these layers. Now, they reported the electrical performance of stabilized lipid monolayers on H-terminated silicon.
Oct 27th, 2011
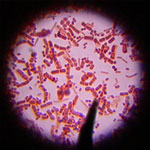 In order to enhance the utilization of nanomaterial in biological systems, it is very important to understand the influence they impart on cellular health and function. Nanomaterials present a research challenge as very little is known about how they behave in relation to micro-organisms, particularly at the cellular and molecular levels. Most of the nanomaterials reported earlier have demonstrated to be efficient antimicrobial agents against virus, bacteria or fungus. There are scarce research reports on the growth-promoting role of nanomaterials especially with respect to microbes. Recent findings, however, have challenged this concept of antimicrobial activity of nanoparticles.
In order to enhance the utilization of nanomaterial in biological systems, it is very important to understand the influence they impart on cellular health and function. Nanomaterials present a research challenge as very little is known about how they behave in relation to micro-organisms, particularly at the cellular and molecular levels. Most of the nanomaterials reported earlier have demonstrated to be efficient antimicrobial agents against virus, bacteria or fungus. There are scarce research reports on the growth-promoting role of nanomaterials especially with respect to microbes. Recent findings, however, have challenged this concept of antimicrobial activity of nanoparticles.
Oct 13th, 2011
 A main difference between central and peripheral nervous system is the lack of regeneration after a neurotrauma, leading to severe and irreversible handicaps. While biomaterials have been developed to aid the regeneration of peripheral nerves, the repair of central nerves such as the optic nerval or nerve cells in the spinal cord remain a major challenge for scientists. The ability to regenerate central nerve cells in the body could reduce the effects of trauma and disease in a dramatic way and nanotechnologies offer promising routes for repair techniques. Scientists have now attempted to rescue retinal ganglion cell death and enhance their regeneration using an electrospun material made of biofunctional nanofibers.
A main difference between central and peripheral nervous system is the lack of regeneration after a neurotrauma, leading to severe and irreversible handicaps. While biomaterials have been developed to aid the regeneration of peripheral nerves, the repair of central nerves such as the optic nerval or nerve cells in the spinal cord remain a major challenge for scientists. The ability to regenerate central nerve cells in the body could reduce the effects of trauma and disease in a dramatic way and nanotechnologies offer promising routes for repair techniques. Scientists have now attempted to rescue retinal ganglion cell death and enhance their regeneration using an electrospun material made of biofunctional nanofibers.
Oct 12th, 2011
 We are experiencing an unprecedented resurgence of interest in herbal healing, and 'herbal renaissance' is happening all over the globe. The Western world has begun to acknowledge the importance of traditional medicines as they symbolize safety in contrast to the allopathic medicines, which tend to produce undesirable side effects and are lacking in curative value. In the realm of medicine, nanotechnology holds enormous promise for benefitting society by potentially reducing the miseries of people suffering from grave illnesses and save a great number of lives. Traditional Oriental medicine would greatly benefit by integrating with the scientific advancements in medical science and diagnostics in concert with nanotechnology. This trinity may usher in a new era of affordable, safe and effective medicinal system.
We are experiencing an unprecedented resurgence of interest in herbal healing, and 'herbal renaissance' is happening all over the globe. The Western world has begun to acknowledge the importance of traditional medicines as they symbolize safety in contrast to the allopathic medicines, which tend to produce undesirable side effects and are lacking in curative value. In the realm of medicine, nanotechnology holds enormous promise for benefitting society by potentially reducing the miseries of people suffering from grave illnesses and save a great number of lives. Traditional Oriental medicine would greatly benefit by integrating with the scientific advancements in medical science and diagnostics in concert with nanotechnology. This trinity may usher in a new era of affordable, safe and effective medicinal system.
Sep 8th, 2011
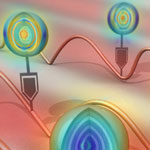 Most molecular probes used in biomedical research require dyes or fluorescence in order to obtain meaningful signals. These probes usually are quite limited with regard to the complexity of what they can image - be it the measurable concentration range or the number of molecules that can be simultaneously detected. This is an issue that is particularly relevant when it comes to track the simultaneous multiple molecular transformations that dictate complicated diseases like cancer. Scientists now have come up with an intriguing new class of molecular probes to solve this problem. They took an existing spectroscopic technique - surface-enhanced Raman scattering (SERS) - and developed a unique class of nanoparticle labels that provide for different responses when excited by laser light.
Most molecular probes used in biomedical research require dyes or fluorescence in order to obtain meaningful signals. These probes usually are quite limited with regard to the complexity of what they can image - be it the measurable concentration range or the number of molecules that can be simultaneously detected. This is an issue that is particularly relevant when it comes to track the simultaneous multiple molecular transformations that dictate complicated diseases like cancer. Scientists now have come up with an intriguing new class of molecular probes to solve this problem. They took an existing spectroscopic technique - surface-enhanced Raman scattering (SERS) - and developed a unique class of nanoparticle labels that provide for different responses when excited by laser light.
Aug 30th, 2011
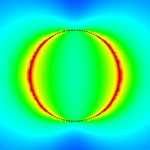 Extracellular signaling molecules are the language that cells use to communicate with each other. These molecules transfer information not only via their chemical compositions but also through the way they are distributed in space and time throughout the cellular environment. With the development of nanosensing techniques, scientists are trying to to eavesdrop on the cellular whisper and they getting closer to deciphering extracellular signaling - an important task in understanding how cells organize themselves, for instance during organ development or immune responses. Now, researchers have reported a novel sensing technique to interrogate extracellular signaling at the subcellular level. They developed a nanoplasmonic resonator array to enhance fluorescent immunoassay signals up to more than one hundred times to enable the first time submicrometer resolution quantitative mapping of endogenous cytokine secretion from an individual cell in nanoscale close to the cell.
Extracellular signaling molecules are the language that cells use to communicate with each other. These molecules transfer information not only via their chemical compositions but also through the way they are distributed in space and time throughout the cellular environment. With the development of nanosensing techniques, scientists are trying to to eavesdrop on the cellular whisper and they getting closer to deciphering extracellular signaling - an important task in understanding how cells organize themselves, for instance during organ development or immune responses. Now, researchers have reported a novel sensing technique to interrogate extracellular signaling at the subcellular level. They developed a nanoplasmonic resonator array to enhance fluorescent immunoassay signals up to more than one hundred times to enable the first time submicrometer resolution quantitative mapping of endogenous cytokine secretion from an individual cell in nanoscale close to the cell.
Aug 15th, 2011
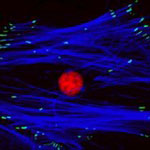 Currently, when adult stem cells are harvested from a patient, they are cultured in the laboratory to increase the initial yield of cells and create a batch of sufficient volume to kick-start the process of cellular regeneration when they are re-introduced back into the patient. The process of culturing is made more difficult by spontaneous stem cell differentiation, where stem cells grown on standard plastic tissue culture surfaces do not expand to create new stem cells but instead create other cells which are of no use in therapy. New findings show that nanoscale patterning is a powerful tool for the non-invasive manipulation of stem cells. Their facile fabrication process employed, a range of thermoplastics that can be processed with exquisite reproducibility down to 5 nm fidelity using injection moulding approaches, offers unique potential for the generation of cell culture platforms for the up-scale of autologous cells for clinical use.
Currently, when adult stem cells are harvested from a patient, they are cultured in the laboratory to increase the initial yield of cells and create a batch of sufficient volume to kick-start the process of cellular regeneration when they are re-introduced back into the patient. The process of culturing is made more difficult by spontaneous stem cell differentiation, where stem cells grown on standard plastic tissue culture surfaces do not expand to create new stem cells but instead create other cells which are of no use in therapy. New findings show that nanoscale patterning is a powerful tool for the non-invasive manipulation of stem cells. Their facile fabrication process employed, a range of thermoplastics that can be processed with exquisite reproducibility down to 5 nm fidelity using injection moulding approaches, offers unique potential for the generation of cell culture platforms for the up-scale of autologous cells for clinical use.
Jul 20th, 2011
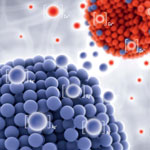 Silicone elastomers are widely used for biomedical applications and products. One major challenge for biomedical applications is to control the ingrowth of silicone-based implants and to avoid bacterial infections on device surfaces. The use of ions from metals like silver and copper is a promising, long-lasting method to achieve such bioactive effects. Researchers have now found a novel effect caused by a combination of copper and silver nanoparticles in silicone. By fabricating bioactive nanocomposite materials that release these ions in specific concentration levels and during a long time, manufacturers can control the bioactive effects of their medical devices or implants.
Silicone elastomers are widely used for biomedical applications and products. One major challenge for biomedical applications is to control the ingrowth of silicone-based implants and to avoid bacterial infections on device surfaces. The use of ions from metals like silver and copper is a promising, long-lasting method to achieve such bioactive effects. Researchers have now found a novel effect caused by a combination of copper and silver nanoparticles in silicone. By fabricating bioactive nanocomposite materials that release these ions in specific concentration levels and during a long time, manufacturers can control the bioactive effects of their medical devices or implants.
Jul 8th, 2011
 Lipids are the main component of the outermost membrane of cells. Their role is to seperate the inner and outer media of the cell and prevent any ionic current between these two media. Because of this last property, lipid layers can be thought of as good ultra-thin insulators that could be used in the development of electronic devices. So far though, because of their inherent instability in air, their use in advanced processes has been limited. This might change, though, since researchers in France have shown the possibility to stabilize by polymerization a lipid monolayer with a thickness of 2.7 nm directly at the surface of H-terminated silicon surface therefore opening a whole new world of possibilities of the use of these layers. Now, they reported the electrical performance of stabilized lipid monolayers on H-terminated silicon.
Lipids are the main component of the outermost membrane of cells. Their role is to seperate the inner and outer media of the cell and prevent any ionic current between these two media. Because of this last property, lipid layers can be thought of as good ultra-thin insulators that could be used in the development of electronic devices. So far though, because of their inherent instability in air, their use in advanced processes has been limited. This might change, though, since researchers in France have shown the possibility to stabilize by polymerization a lipid monolayer with a thickness of 2.7 nm directly at the surface of H-terminated silicon surface therefore opening a whole new world of possibilities of the use of these layers. Now, they reported the electrical performance of stabilized lipid monolayers on H-terminated silicon.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





