| Posted: Nov 17, 2015 |
3-D mapping a new drug-delivery tool
(Nanowerk News) Cubosomes are small biological 'capsules' that can deliver molecules of nutrients or drugs with high efficiency. They have a highly symmetrical interior made of tiny cubes of assembled fat molecules similar to the ones in cell membranes. This also means that cubosomes are safe to use in living organisms. Such features have triggered great interest in the pharmaceutical and food industry, who seek to exploit the structure of cubosomes for the controlled release of molecules, improving the delivery of nutrients and drugs.
|
|
Ecole Polytechnique Fédérale de Lausanne (EPFL) scientists, working with Nestlé, have now been able to study the 3D structure of cubosomes in detail for the first time. Published in Nature Communications ("Direct visualization of dispersed lipid bicontinuous cubic phases by cryo-electron tomography"), the breakthrough can help and promote the use of cubosomes in medicine and food science.
|
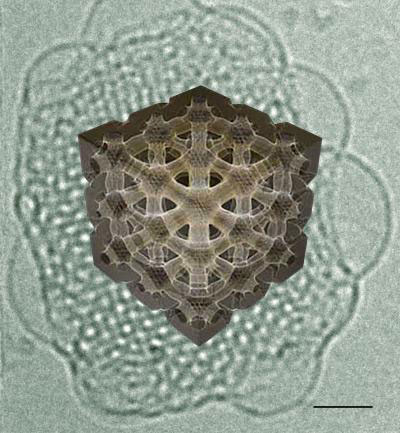 |
| 3-D reconstruction of one of the 2 water channels of an inverse bicontinuous cubic phase obtained from cryo-electron tomography experiments. Scale bar: 40 nm. (Image: Davide Demurtas/EPFL)
|
|
Molecules of a drug or a nutrient contained inside a cubosome can move by using the numerous tiny channels that make up its structure's interior. The pharmaceutical industry already uses a similar system for drug delivery: the liposomes, which are also made of fats but in the shape of a sphere. Their intricate internal channels give cubosomes a very high internal surface, which offers great potential for the controlled delivery of nutrients and drugs.
|
|
In short, the properties of cubosomes, like other lipid-based delivery vessels, depend on their particular structures. The problem is that cubosomes are self-assembled, occurring 'spontaneously' after putting together the right ingredients (generally fats and a detergent) under the right conditions. This means that scientists have limited control over their final structure, which makes it hard to optimize their design. In addition, it is very difficult to 'see' the interior of a cubosome and map out the various arrangements of its channels.
|
|
Davide Demurtas and Cécile Hébert from EPFL's Interdisciplinary Centre for Electron Microscopy (CIME), working with Laurent Sagalowicz at the Nestlé Research Center in Lausanne, have now uncovered the interior 3D structure of cubosomes, and have successfully matched their real-life findings to computer simulations.
|
|
The researchers used a microscopy technique called 'cryo-electron tomography' (CET). Their method involves embedding cubosomes in a type of 'glass' ice that does not form crystals, which would damage the cubosomes. The samples are kept at -170oC. The microscope then takes photographs while tilting the cubosome at different angles. The technique, which was carried out at CIME, can reconstruct the three-dimensional information to create images of the cubosomes in their native state and with unprecedented detail.
|
|
"This method allows us to get information about everything, both the inside and outside of the cubosomes," says Cécile Hébert. "Because the CET microscope distinguishes the different densities between cubosome and ice, it is very sensitive and precise."
|
|
The CET images clearly showed the internal cubic structure, as well as the internal 3D organization of the channels. The researchers also compared the images to the prevailing mathematical models used to make computer simulations of the interface between the interior and exterior. The real-life data successfully matched the theory.
|
|
"With this approach we can now forge a new understanding of the structure of the cubosomes' interior," says Davide Demurtas. The success is expected to make the study and design of cubosomes with controlled macroscopic properties (e.g. controlled release) easier.
|

