| Posted: Jan 25, 2016 |
Acoustic tweezers provide much needed pluck for 3D bioprinting
(Nanowerk News) Researchers, including Carnegie Mellon University President Subra Suresh and collaborators Tony Jun Huang from the Pennsylvania State University and Ming Dao from MIT, have demonstrated that acoustic tweezers can be used to non-invasively move and manipulate single cells along three dimensions, providing a promising new method for 3-D bioprinting. Their findings are published in this week’s issue of the Proceedings of the National Academy of Sciences ("Three-dimensional manipulation of single cells using surface acoustic waves").
|
|
Multicellular structures within living things are complex and delicate, which makes recreating these structures a daunting task. For example, the human heart contains more than 2 billion muscle cells. Each of these cells must properly interact with one another and with their environment to ensure that the heart functions properly. If those cells aren’t placed correctly, or are damaged, it could potentially result in any of a variety of heart conditions.
|
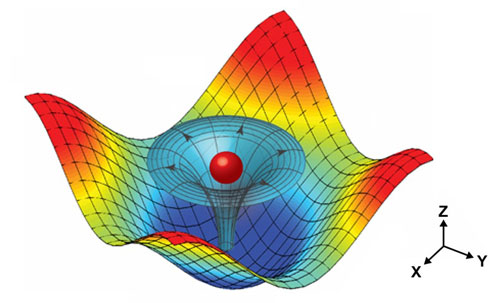 |
| Illustration of a particle trapped by the 3-D trapping node created by two superimposed, orthogonal, standing surface acoustic waves and the induced acoustic streaming.
|
|
3-D bioprinting is a promising way to recreate the complex, multicellular architecture of biological tissues. Researchers have been using a combination of approaches, but have yet to develop a single method that has the high level of precision, versatility, multiple dimensionality and single cell resolution needed to form complex multicellular structures while maintaining cell viability, integrity and function.
|
|
“The results presented in this paper provide a unique pathway to manipulate biological cells, accurately and in three dimensions, without the need for any invasive contact, tagging or biochemical labeling,” Suresh said. “This approach could lead to new possibilities for research and applications in such areas as regenerative medicine, neuroscience, tissue engineering, bio-manufacturing and cancer metastasis.”
|
|
The Carnegie Mellon, Penn State and MIT team has been at the forefront of developing acoustic tweezers technology, a technique that uses sound waves to trap and manipulate single cells. Acoustic tweezers have been used to separate, align, pattern and transport single cells, and are renowned for their ability to gently manipulate cells without causing any cellular damage. In order to make the technology functional for 3-D printing, the researchers needed to prove that it could capture and manipulate cells along all three dimensions.
|
|
The researchers’ most recent version of acoustic tweezers involves a microfluidic device that uses acoustic wave generators to produce sound waves along the edges of the device. The device’s design allowed researchers to manipulate where the waves would meet along each of the three axes. At these meeting points, the waves formed a 3-D trapping node that captured individual cells. The researchers could then further manipulate the acoustic waves to move and place cells.
|
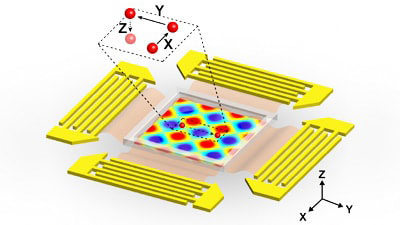 |
| Illustration of the planar surface acoustic wave generators, used to generate volumetric nodes, surrounding the microfluidic experimental area. The inset indicates a single particle within a “3-D trapping node,” which is independently manipulated along the x, y or z axes.
|
|
“This approach could lead to new possibilities for research and applications in such areas as regenerative medicine, neuroscience, tissue engineering, bio-manufacturing and cancer metastasis.” — Subra Suresh
|
|
To demonstrate how their acoustic tweezers technique could be used for live cell printing, the researchers used the microfluidic device to pick up cells and deposit them in a pre-selected pattern. The device displayed an elegant level of control over cell spacing and geometry. This indicated to researchers that the device has the potential to effectively create 3-D tissue-like structures, including those with complex geometries.
|


