| Posted: Mar 18, 2016 |
Scientists develop new technique for imaging cells and tissues under the skin
(Nanowerk News) Scientists have many tools at their disposal for looking at preserved tissue under a microscope in incredible detail, or peering into the living body at lower resolution. What they haven't had is a way to do both: create a three-dimensional real-time image of individual cells or even molecules in a living animal.
|
|
Now, Stanford scientists have provided the first glimpse under the skin of a living animal, showing intricate real-time details in three dimensions of the lymph and blood vessels.
|
|
The technique, called MOZART (for MOlecular imaging and characteriZation of tissue noninvasively At cellular ResoluTion), could one day allow scientists to detect tumors in the skin, colon or esophagus, or even to see the abnormal blood vessels that appear in the earliest stages of macular degeneration - a leading cause of blindness.
|
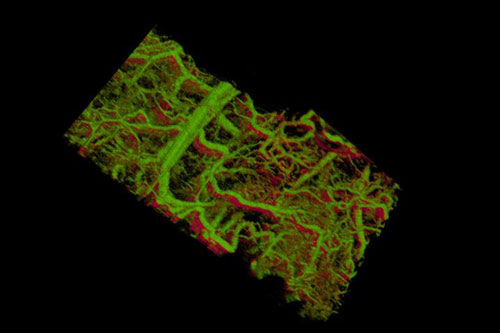 |
| Gold nanorods within the blood vessels of a mouse ear appear green. The lower right shows vessels within a tumor that lies under the skin. (Image: de la Zerda lab)
|
|
"We've been trying to look into the living body and see information at the level of the single cell," said Adam de la Zerda, an assistant professor of structural biology at Stanford and senior author on the paper. "Until now there has been no way do that."
|
|
De la Zerda, who is also a member of Stanford Bio-X, said the technique could allow doctors to monitor how an otherwise invisible tumor under the skin is responding to treatment, or to understand how individual cells break free from a tumor and travel to distant sites.
|
|
Going for gold
|
|
A technique exists for peeking into a live tissue several millimeters under the skin, revealing a landscape of cells, tissues and vessels. But that technique, called optical coherence tomography, or OCT, isn't sensitive or specific enough to see the individual cells or the molecules that the cells are producing, which is what interests de la Zerda.
|
|
A major issue has been finding a way of differentiating between cells or tissues; for example, picking out the cancerous cells beginning to multiply within an overall healthy tissue. In other forms of microscopy, scientists have created tags that latch onto molecules or structures of interest to illuminate those structures and provide a detailed view of where they are in the cell or body.
|
|
No such beacons existed for OCT, though de la Zerda knew that tiny particles called gold nanorods had some of the properties he was looking for. The problem was that the commercially available nanorods didn't produce nearly enough signal to be detected in a tissue.
|
|
What the team needed were nanorods, but big ones. Nanorods are analogous to organ pipes, said graduate student Elliott SoRelle, because longer pipes vibrate at lower frequencies, creating a deep, low sound. Likewise, longer nanorods vibrate at lower frequencies, or wavelengths, of light. Those vibrations scatter the light, which the microscope detects.
|
|
If all the other tissues are vibrating in a white noise of higher frequencies, longer nanorods would stand out like low organ notes amidst a room of babble.
|
|
SoRelle's challenge was to manufacture longer nanorods that were nontoxic, stable and very bright, which turned out to be a lot to ask. "My background was biochemistry, and this turned out to be a problem of materials science and surface chemistry," said SoRelle, who was co-first author on the paper. He can now make nontoxic nanorods in various sizes that all vibrate at unique and identifiable frequencies.
|
|
Eliminating noise
|
|
The next challenge was filtering out the nanorods' frequency from the surrounding tissue.
|
|
To do that, electrical engineering graduate student and Bowes Bio-X Fellow Orly Liba developed computer algorithms that could separate out the frequencies of light scattered by nanorods of various lengths and differentiate those from surrounding tissue.
|
|
With SoRelle's large nanorods and Liba's sensitive algorithms, de la Zerda and his team had solved the initial problem of detecting specific structures in three-dimensional images of living tissues. The resulting three-dimensional, high-resolution images were so big - on the order of gigapixels - that the team needed to develop additional algorithms for analyzing and storing such large images.
|
|
The team tested their technology in the ear of a living mouse, where they were able to watch as the nanorods were taken up into the lymph system and transported through a network of valves. They were able to distinguish between two different size nanorods that resonated at different wavelengths in separate lymph vessels, and they could distinguish between those two nanorods in the lymph system and the blood vessels. In one study, they could watch individual valves within the lymph vessels open and close to control the flow of fluid in a single direction.
|
|
"Nobody has shown that level of detail before," said Liba, who was co-first author on the paper.
|
|
Impossible goal
|
|
This detailed imaging was de la Zerda's initial goal when he started his lab in 2012, though he was frequently told it would be impossible. "I'm in a small department, but with very accomplished faculty," he said. "One faculty member told me his own life story of taking big risks and that encouraged me. I thought it would be really fun to see if we can make it work and see cells talking to each other in real time."
|
|
His gamble got off the ground primarily with a seed grant from Stanford Bio-X, which supports early-stage interdisciplinary research. "That grant allowed us to take a big risk in a direction that was completely unproven," de la Zerda said.
|
|
Having shown that the gold nanorods can be seen in living tissue, the next step is to show that those nanorods can bind to specific kinds of cells, like skin cancer or abnormal vessels in early stage macular degeneration. Then, the technique could be used to learn more about how those diseases progress at the molecular level and also evaluate treatments in individual patients, something that previously hadn't been possible.
|

