| Posted: Sep 28, 2016 |
Researchers suggest nanoscale electronic motion sensor as DNA sequencer
(Nanowerk News) Researchers from the National Institute of Standards and Technology (NIST) and collaborators have proposed a design for the first DNA sequencer based on an electronic nanosensor that can detect tiny motions as small as a single atom.
|
|
The proposed device—a type of capacitor, which stores electric charge—is a tiny ribbon of molybdenum disulfide suspended over a metal electrode and immersed in water. The ribbon is 15.5 nanometers (nm, billionths of a meter) long and 4.5 nm wide. Single-stranded DNA, containing a chain of bases (bits of genetic code), is threaded through a hole 2.5 nm wide in the thin ribbon. The ribbon flexes only when a DNA base pairs up with and then separates from a complementary base affixed to the hole. The membrane motion is detected as an electrical signal.
|
|
As described in a new paper (ACS Nano, "A MoS2-based capacitive displacement sensor for DNA sequencing"), the NIST team made numerical simulations and theoretical estimates to show the membrane would be 79 to 86 percent accurate in identifying DNA bases in a single measurement at speeds up to about 70 million bases per second. Integrated circuits would detect and measure electrical signals and identify bases. The results suggest such a device could be a fast, accurate and cost-effective DNA sequencer, according to the paper.
|
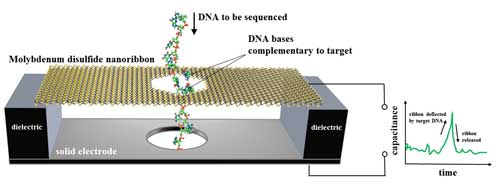 |
| NIST's proposed design for a DNA sequencer based on an electronic motion sensor. A nanoscale ribbon of molybdenum disulfide is suspended over a metal electrode and immersed in water. Single-stranded DNA, containing a chain of bases (bits of genetic code), is threaded through a hole in the ribbon, which flexes only when a DNA base pairs up with and then separates from a complementary base affixed to the hole. The membrane motion is detected as an electrical signal. Numerical simulations and theoretical estimates show the membrane would be 79 to 86 percent accurate in identifying DNA bases in a single measurement at speeds up to about 70 million bases per second. (Image: NIST) (click on image to enlarge)
|
|
Conventional sequencing, developed in the 1970s, involves separating, copying, labeling and reassembling pieces of DNA to read the genetic information. Newer methods include automated sequencing of many DNA fragments at once—still costly—and novel “nanopore sequencing” concepts. For example, the same NIST group recently demonstrated the idea of sequencing DNA by passing it through a graphene nanopore, and measuring how graphene’s electronic properties respond to strain.
|
|
The latest NIST proposal relies on a thin film of molybdenum disulfide—a stable, layered material that conducts electricity and is often used as a lubricant. Among other advantages, this material does not stick to DNA, which can be a problem with graphene. The NIST team suggests the method might even work without a nanopore—a simpler design—by passing DNA across the edge of the membrane.
|
|
“This approach potentially solves the issue with DNA sticking to graphene if inserted improperly, because this approach does not use graphene, period,” NIST theorist and lead author Alex Smolyanitsky said. “Another major difference is that instead of relying on the properties of graphene or any particular material used, we read motions electrically in an easier way by forming a capacitor. This makes any electrically conductive membrane suitable for the application.”
|
|
Nanomaterials expert Boris Yakobson of Rice University, a co-author on the paper, suggested the capacitor idea. Computational support was provided by the University of Groningen in the Netherlands.
|
|
DNA has four bases. For the simulations, cytosine (C), which naturally pairs up with guanine (G), is attached to the inside of the pore. When a piece of DNA passes through the pore, any G in the strand temporarily attaches to the embedded C, pulling on the nanoribbon and signaling the electrode. The DNA sequence is determined by measuring how and when electrical blips vary over time. To detect all four bases, four nanoribbons, each with a different base attached to the pore, could be stacked vertically to create an integrated DNA sensor.
|
|
The molybdenum disulfide ribbon is flexible enough to deform measurably in response to the forces required to break up a DNA pair, but rigid enough to have less ongoing, meaningless movement than graphene, potentially reducing unwanted noise in the sequencing signals. The deflection of the ribbon is exceedingly small, on the order of one angstrom, the size of a hydrogen atom. Its pulling force is on the order of 50 piconewtons, or trillionths of a newton, enough to break up the delicate chemical bonds between DNA bases.
|
|
Researchers estimated how the device would perform in an integrated circuit and found the peak currents through the capacitor were measurable (50 to 70 picoamperes), even for the small nanoribbons studied. The current peaks are expected to be even larger in physical systems. The device size could be tweaked to make it even easier to measure sequencing signals.
|
|
The NIST authors hope to build a physical version of the device in the future. For practical applications, the chip-sized DNA sequencing microfluidic technology might be combined with electronics into a single device small enough to be handheld.
|

