| Posted: Jul 25, 2017 |
Carbon nanotubes stand at attention
(Nanowerk News) Just as members of a marching band align themselves for a performance, carbon nanotubes create a similar configuration.
|
|
Lawrence Livermore National Laboratory (LLNL) scientists recently used synchrotron X-ray scattering to fully capture the hierarchical structure in self-organized carbon nanotube materials from the atomic to micrometer scale. Their work, recently published in the June issue of ACS Nano ("Quantifying the Hierarchical Order in Self-Aligned Carbon Nanotubes from Atomic to Micrometer Scale"), is the first to continuously map the structural order of nanotube ensembles across four orders of magnitude in length scale, all while employing a single technique.
|
|
Complex hierarchical structures made from synthetic nanocarbon allotropes such as nanotubes and graphene promise to transform countless applications in structural composites, nanoelectronics, energy storage, filtration and separation. Just as the arrangement of atoms and defects classically govern a material's function, in a similar way the order and alignment of nanoscale building blocks within a larger ensemble strongly influence the material's macroscale performance.
|
|
A lack of comprehensive, multiscale structural characterization has been a crucial bottleneck to progress in application-targeted synthesis of hierarchical nanomaterials.
|
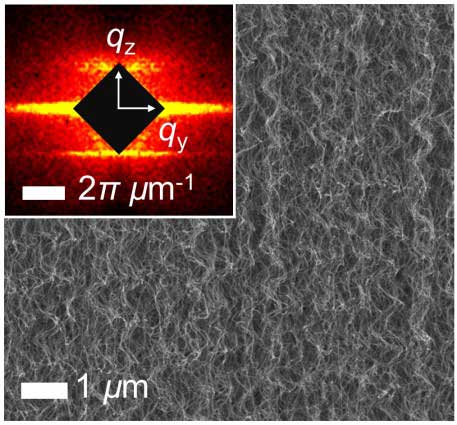 |
| Microscale corrugated structure of a vertically aligned CNT 'forest' revealed by electron microscopy (real space) and corresponding inset soft X-ray scattering data (inverse space). (Image: LLNL)
|
|
"We were interested in describing the entire structure of aligned carbon nanotube 'forests' across dramatically different length scales, which typically cannot be done using just one technique, such as conventional microscopy or spectroscopy," noted Eric Meshot, LLNL scientist and lead author on the study. "X-ray scattering is powerful because the addressable feature size is widely tunable simply based on the incoming X-ray energy and where you place your detector to collect the outgoing X-rays."
|
|
This approach enabled team members to draw correlations between adjacent length scales, which revealed that the packing density of nanotubes ultimately influences alignment at every length scale. Notably, the researchers forged new ground by using soft (low-energy) X-rays to resolve microscale structural patterns that can emerge along the nanotube growth direction. Surprisingly, they found that these carbon nanotube materials may form vertical corrugations with high microscale order despite having low nanoscale order.
|
|
The impact of this study goes beyond fundamental understanding of structure. The LLNL team has used X-ray scattering as a workhorse capability for evaluating the structure-performance relationship in aligned carbon nanotube membranes toward building breathable garments that protect against biological threats. "Structural characteristics like pore size distribution, pore density and tortuosity dictate membrane transport performance and can be quantified easily with X-ray methods," explained Francesco Fornasiero, LLNL scientist and the principal investigator on the project.
|
|
For this work, the team leveraged a tight collaboration with the Advanced Light Source (ALS) and the Molecular Foundry. "We would like to see more of this type of 'cross-pollination' between DOE facilities so that our users can fully exploit cutting-edge structural characterization at the ALS to inform nanostructure synthesis at the Foundry," said Teyve Kuykendall, a principal scientific engineer at the Molecular Foundry and coauthor on the study.
|
|
"We are excited about moving forward to explore how we can use X-ray scattering tools to decipher in real-time material structure as a function of length scale, time and chemistry all together," added Cheng Wang, a staff scientist at the ALS and coauthor on this work. This array of information would be significant for establishing multiscale structure-property relationships toward application-oriented design and manufacturing.
|

