| Posted: Aug 23, 2017 |
Self-assembling protein complexes could provide scaffolding for nanostructures
(Nanowerk News) When hemoglobin undergoes just one mutation, these protein complexes stick to one another, stacking like Lego blocks to form long, stiff filaments. These filaments, in turn, elongate the red blood cells found in sickle-cell disease. For over 50 years, this has been the only known textbook example in which a mutation causes such filaments to form.
|
|
According to Dr. Emmanuel Levy and his group in the Weizmann Institute of Science's Structural Biology Department, Lego-like assemblies should have formed relatively frequently during evolution. Could this assembly method be common, or even easy to reproduce? Their answer, which was recently published in Nature ("Proteins evolve on the edge of supramolecular self-assembly"), may have implications for both biological research and nanoscience.
|
|
Hemoglobin and a fair number of other protein complexes are symmetric: made of identical units. And since identical units are produced from the same gene, each genetic mutation is repeated multiple times in the complex. Mutations that create sticky patches and are repeated on opposite sides of the complex can induce the proteins to stack into long protein fibers. Unlike amyloid-like protein fibers, the complexes in these stacks do not change shape or unfold in order to assemble.
|
|
The stickiness occurs because the mutation substitutes an amino acid that is normally hydrophilic -- "water-loving" -- with one that is hydrophobic -- "water-hating." In the watery environment in which proteins move, the hydrophobic regions on those proteins prefer to interact with one another, like foam bubbles in water.
|
|
In their experiments, Levy and his group, including Hector Garcia-Seisdedos, Charly Empereur-Mot (who is now at Conservatoire National des Arts et Métiers in Paris) and Nadav Elad of the Weizmann Institute's Chemical Research Support Department, began with an ultra-symmetric protein complex made up of eight identical units. They followed just one rule for mutating the proteins: Switch a hydrophilic amino acid with a hydrophobic, "sticky," one.
|
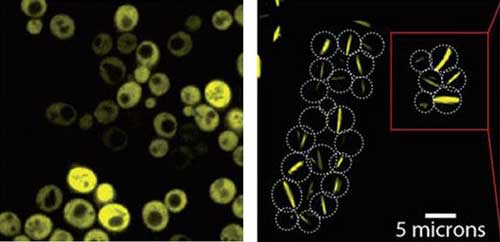 |
| Yeast cells producing a bacterial symmetric protein complex with eight units. When it is not mutated (left), the complex diffuses freely inside the cell, but a single mutation (right) triggers its assembly into long filaments. (Image: Weizmann Institute of Science)
|
|
The team initially created proteins with three mutations to two different sticky amino acids and observed Lego-like self-assembly in both cases. Investigating further, the team experimented with each mutation individually and found that one was capable, on its own, of producing the long filaments.
|
|
So, are mutations that only do one thing -- increase the stickiness of the protein's surface -- likely to induce Lego-like self-assembly? The researchers mutated 11 additional proteins known to form symmetric complexes - creating 73 different mutations in all -- and produced them in baker's yeast cells, adding a fluorescent protein "label" to enable their visualization. In 30 of these variations, the researchers observed behavior that suggested self-assembly: Around half of these had stacked into long filaments, while the other half were bunched together in a more amorphous way, forming "foci."
|
|
If the researchers reproduced the phenomenon of sickle-cell filaments so easily in the lab, why is it not seen more in biomedical research? Levy proposes two answers: Firstly, the team revealed that naturally symmetric proteins evolved to have extra hydrophilic amino acids on their surfaces, thus minimizing the risk of self-assembly. Secondly, says Levy, researchers probably see more Lego assemblies than they think: "Now that researchers know they can evolve so readily, they may look at foci more carefully and see many more biologically relevant Lego-like assemblies."
|
|
"Also," he adds, "the filaments are produced so easily in the yeast, they could be good candidates for the scaffolding of nanostructures. Our study was unique in that it did not require complex computational design, nor did we have to scan thousands of mutations to find the one we wanted. We simply started with an existing structure and found a simple strategy to induce the assembly of filaments."
|

