| Posted: Dec 21, 2017 |
Nanofractionation platform with parallel mass spec to ID cytochrome CYP1A2 inhibitors
(Nanowerk News) A new (and freely available) original research article by Barbara M. Zietek et al., now available ahead-of-print at SLAS Discovery Online ("Nanofractionation Platform with Parallel Mass Spectrometry for Identification of CYP1A2 Inhibitors in Metabolic Mixtures"), presents a fast, robust and accurate methodology for correlating compound identity to CYP1A2 potency of inhibitors in metabolic mixtures.
|
|
The methodology is centered around an at-line nanofractionation platform in which a metabolic mixture is chromatographically separated followed by parallel on-line mass spectrometric (MS) analysis and at-line nanofractionation on high-density microtiter well plates that are then directly exposed to a bioassay.
|
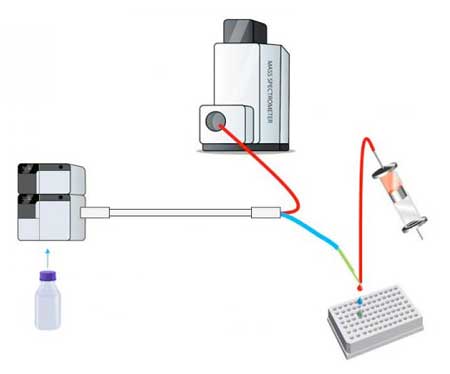 |
| Generation of Phase I metabolic mixture of a drug is followed by chromatographic separation of the metabolites. A split directs eluent to MS for compound identification and to nanofractionation on a 384-well plate for bioassaying. Finally, a bioassay chromatogram is reconstructed and comparison with parallel obtained MS data is performed. (Image: Jeroen Kool)
|
|
Correlation of reconstructed bioactivity chromatograms with MS data allows direct identification of compounds with inhibitory properties toward CYP1A2 enzymes, indicating CYP-involved drug-drug interactions. In a similar fashion, the methodology can be implemented in assaying metabolic mixtures for other CYPs relevant in drug discovery. Next to traditional Met-ID metabolic profiling, the approach presented in this new article provides direct profiles of the relative contribution of each metabolite to the assayed activity, in this case CYP1A2 inhibition.
|
|
Drug-drug interactions caused by inhibition of CYP450 enzymes (or CYPs) can lead to serious adverse reactions in patients. These inhibitory properties can be exhibited by parent compounds and by their metabolites. Together, the properties of one or more metabolites can outweigh the benefits of a drug candidate, thus leading to its failure in the drug development phase or removal from the pharmaceutical market after introduction.
|
|
In recent years, increases in drug (candidate) attrition is especially seen in the later stages of drug development. This process can be improved by implementing new screening methods that rapidly assess the biological/biochemical properties and correlate them directly to compound identities. This can rapidly deliver profiles of pharmacokinetic properties of a drug and its metabolites together with their biological properties and can be achieved by the methodology presented in this article.
|

