| Mar 03, 2021 |
Researchers demonstrate highly efficient 3D nano-imaging with XFEL and machine learning
(Nanowerk News) It is millions of trillions of times brighter than the sunlight and a whopping 1,000 trillionth of a second, appropriately called the instantaneous light. It is the X-ray Free Electron Laser (XFEL) light that opens a new scientific paradigm.
|
|
Combining it with AI, an international research team has succeeded in filming and restoring the 3D structure of nanoparticles that share structural similarities with viruses (ACS Nano, "High-Throughput 3D Ensemble Characterization of Individual Core–Shell Nanoparticles with X-ray Free Electron Laser Single-Particle Imaging").
|
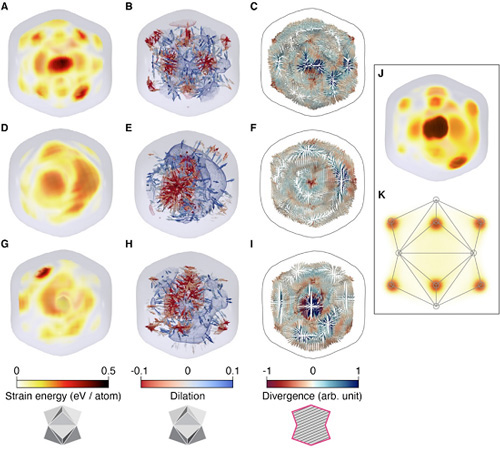 |
| Elastic strain analysis. Three-dimensional map of strain energy and strain field of the nanoparticles (NPs). (A) 3D distribution of strain energy accumulated on the type-I NP. Higher strain energy is found at the position corresponding to the vertices of the trisoctahedron. (B) Strain field in 3D shown using arrows indicating the direction for higher strain energy. The strain energy distribution is overlaid using a contour map with the regions of local expansion in blue and compression in red. (C) Strain visualized using color-scaled arrows exhibiting locally contracted and relaxed regions at the central plane of the NP. Similar plots of 3D strain energy distribution (D, G), 3D strain field (E, H), and detailed strain field on a plane (F, I) obtained for type-II and type-III NPs, respectively. (J) Strain energy distribution of the type-I NC calculated from the MD. (K) Comparison of strain energy distribution expected for the ideal TOH Au NC obtained from MD simulations. (© ACS Nano)
|
|
With the fear of a new pandemic growing around the world due to COVID-19, this discovery is attracting the attention among academic circles for imaging the structure of the virus with both high accuracy and speed.
|
|
An international team of researchers from POSTECH, National University of Singapore (NUS), KAIST, GIST, and IBS have successfully analyzed the structural heterogeneities in 3D structures of nanoparticles by irradiating thousands of nanoparticles per hour using the XFEL at Pohang Accelerator Laboratory (PAL) in Korea and restoring 3D multi-models through machine learning. The research team led by Professor Changyong Song and Ph.D. candidate Do Hyung Cho of Department of Physics at POSTECH has driven the international research collaboration to realize it.
|
|
Nanoparticles have a peculiar function that may not be available from native bulk materials, and one can control their physical and chemical properties by designing 3D structures and compositions of constituting elements.
|
|
The commonality between nanoparticles and viruses is that they exist in the form of independent particles, rather than in crystal-regular, periodic arrangements, and, as such, their structures are not uniform at the nanometer level. To precisely understand their structures, it is necessary to statistically analyze the structure of individual particles using the whole ensemble distribution of structures from thousands to hundreds of thousands of specimens.
|
|
However, electron microscopes often fall short of providing enough penetration to limit the size of the sample to be probed; conventional X-rays may damage the sample by X-ray radiation itself, making it difficult to obtain sufficient resolution.
|
|
The research team overcame the practical limitations of the conventional method by using the X-ray free electron laser and the machine learning method to observe the statistical distribution of the 3D structure of thousands of nanoparticles at the nanometer level. As a result, 3D structures of nanoparticles having a size of 300 nm were obtained with a resolution better than 20 nm.
|
|
This achievement was particularly significant for restoring the 3D structure of thousands of nanoparticles using machine learning. Since conventional single-particle imaging techniques often assume an identical 3D structure of the specimens, it was difficult to restore the structure in actual experimental data where the sample structure is not homogeneous.
|
|
However, with the introduction of the multi-model this time, the researchers succeeded in restoring the representative 3D structures. This research has enabled the classification of nanoparticles into four major shapes, and confirmed that about 40% of them had similar structures.
|
|
In addition, through quantitative analysis of the restored 3D structure, the international research collaboration team also uncovered the internal elastic strain distribution accompanied by the characteristic polyhedron structure of the nanoparticles and the inhomogeneous density distribution.
|
|
"These findings enable the observation of 3D structure of noncrystalline viral specimens with inhomogeneously distributed internal molecules," explained Professor Changyong Song of POSTECH. "Adding the 3D image restoration algorithm to this through machine learning shows promise to be applicable to studies of macromolecule structures or viruses in living organisms."
|

