| Posted: March 26, 2010 |
Lost in a crowd |
|
(Nanowerk News) One of the biggest challenges in simulating biological processes is developing mathematical models that accurately reflect the dynamics of real molecules. Koichi Takahashi, of the RIKEN Advanced Science Institute in Yokohama, is in a position to know—as a founding member of the E-Cell Project, he and his colleagues have spent the last 14 years working towards development of a comprehensive simulation of a whole functioning cell.
|
|
Takahashi’s work has focused on dynamic behavior of populations of signaling factors. Such analyses are typically based on ‘mean-field’ descriptions, which assume an essentially averaged distribution of molecules throughout the cellular volume. This shortcut overlooks a lot of real-world complexity, but is made necessary by the excessive computational demands of more accurate models.
|
|
Things have now changed, thanks to a breakthrough from Pieter Rein ten Wolde at AMOLF in the Netherlands, whose team developed an algorithm called Green’s Function Reaction Dynamics (GFRD) that enables sophisticated single-particle-level simulations with considerably reduced computing power ("Green's Function Reaction Dynamics: a new approach to simulate biochemical networks at the particle level and in time and space"). Together, Takahashi and ten Wolde developed a faster and more accurate version of GFRD (Fig. 1), and applied it to mitogen-activated protein kinase (MAPK) cascades, a type of signaling pathway associated with diverse functions in the eukaryotic cell ("Spatio-temporal correlations can drastically change the response of a MAPK pathway").
|
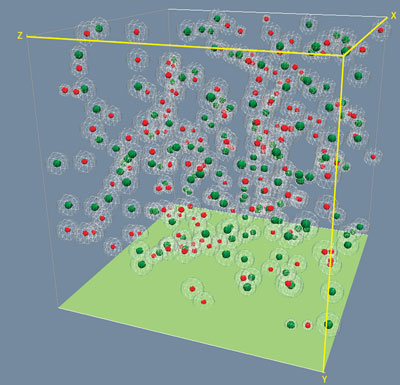 |
| Figure 1: A three-dimensional representation of simulated associations of different molecules of a cell, based on the enhanced GFRD algorithm. (Image: Koichi Takahashi)
|
|
MAPK signaling is a multi-stage process; at each step, one kinase enzyme activates another, downstream kinase through two sequential chemical modifications. Predictions from mean-field-based simulations of MAPK dynamics have depended heavily on whether the enzyme is assumed to act ‘processively’, introducing both modifications without releasing the substrate, or ‘distributively’, releasing the substrate after each modification.
|
|
However, Takahashi and ten Wolde found strikingly different results with their model, which introduces the possibility of re-binding—a scenario in which substrates receive their second modification from the same enzyme that introduced the first. For example, slow diffusion of enzymes and substrate within the cytosol could lead to much more rapid overall activation kinetics by keeping substrates within easy reach of their upstream kinases. “‘Walking slower’ can speed up the response of the system,” says Takahashi. This is the opposite of mean-field model predictions, and essentially erases the distinction between distributive and processive in this context.
|
|
These data demonstrate the importance of increasing complexity in models, a direction in which the authors are continuing to move. “We are extending our method so that we can also include structures such as membranes and organelles,” says Takahashi. “We are also working to make the method even more high-performance, because that will enable us to represent intracellular molecular crowding.”
|

