| Posted: May 14, 2010 |
Fishing for new drug targets with molecule-sized bait |
|
(Nanowerk News) UCLA researchers and their collaborators have developed a method that could open the door for investigations into the function of half of all proteins in the human body.
|
|
The research team has demonstrated nanoscale control over molecules, allowing for the precise study of interactions between proteins and small molecules. Their new technique, in which molecules are used as bait to capture and study large biomolecules, could lead to a new generation of psychiatric medications.
|
|
In a paper published last month in the journal ACS Chemical Neuroscience ("Native Serotonin Membrane Receptors Recognize 5-Hydroxytryptophan-Functionalized Substrates: Enabling Small-Molecule Recognition"), an interdisciplinary team of researchers from UCLA and the Pennsylvania State University (PSU) report on their investigation of the interactions between large biomolecules, which include DNA and proteins, and small molecules, which include hormones and neurotransmitters such as serotonin.
|
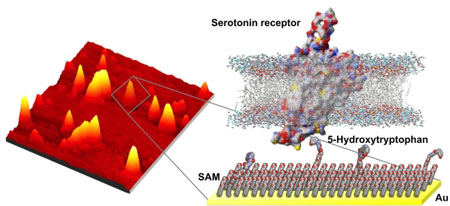 |
| (Left) Atomic force microscopy image of serotonin precursor-modified surface with captured serotonin receptor-containing nanovesicles. (Right) Illustration of the molecular structures of the surface chemistry and the relative size differences between the "bait" (5-hydroxytryptophan) and the membrane-associated serotonin receptors selectively captured by these surfaces.
|
|
The research team, led by Anne Andrews, professor of psychiatry and a researcher at both the Semel Institute for Neuroscience and Human Behavior at UCLA and UCLA's California NanoSystems Institute (CNSI), is studying these interactions to identify a new generation of targets, or key molecules that correspond to specific diseases or conditions.
|
|
Interactions between large biomolecules and small molecules are ubiquitous in nature; they are the method for communication within and between cells. But these interactions have proven difficult to isolate in a laboratory setting. Increased understanding of these interactions is vital for the development of new medications for psychiatric disorders, the researchers say.
|
|
"Currently, little is known about which targets apply to specific diseases," Andrews said. "Pharmaceutical companies are very good at designing medications once they have a target to go after; my group is working on providing them with targets."
|
|
Up to this point, drug development for psychiatric disorders such as depression has been a trial-and-error process in which pharmaceutical companies refine new drugs based on a few existing drugs that were discovered accidentally. Andrews said she hopes that her team's research will lead to more effective treatments, because current depression medications only work for 30 to 50 percent of the population.
|
|
Nanoscale control is the key to the UCLA–Penn State team's findings. Their breakthrough capitalizes on work by the research group of co-author Paul Weiss on patterning self-assembled monolayers (SAMs), single layers of molecules that orient themselves on flat surfaces. Weiss, a distinguished professor of chemistry and biochemistry who holds UCLA's Fred Kavli Chair in Nanosystems Sciences, and others discovered that SAMs don't actually form perfect surfaces. They contain defects, which can in turn be used to isolate single molecules.
|
|
"Currently we are able to space defects out over a surface. We then use these defects to control the placement and environment of the individual functional molecules," said Weiss, who is also director of the CNSI.
|
|
Even spacing is important because the UCLA–Penn State team placed serotonin, a small molecule, in defects to act as bait to capture and study large molecules. If the defects are not widely spaced, there is not enough space between serotonin molecules for each to capture a large molecule.
|
|
Large biomolecule and small molecule interactions have proved notoriously difficult to study using previous methods. When the SAM fishing pole baited with serotonin captures a large molecule, the research team is able to study the interactions in a way that replicates the molecules' natural interactions.
|

