| Posted: July 8, 2010 |
New microscope captures amazing images of embryonic cell development |
|
(Nanowerk News) To understand the complex stages of an embryo, you cannot simply watch it grow. However, it's difficult to accurately observe such fine, complex and high-speed details, even under the microscope. New technical feats now make it possible to capture fruit fly development on video, as well as clear images on how a zebrafish's eyes and midbrain are formed. The German-US team that perfected the method has presented its work in Nature Methods ("Fast, high-contrast imaging of animal development with scanned light sheet–based structured-illumination microscopy").
|
|
The transformation of a cell into a complete animal is a very complex, yet fast process. The original cell divides into tens of thousands of new ones very quickly; they move within the embryo and develop into different organs.
|
|
Thanks to automated image processing and data analysis, scientists are able to follow the behaviour of cells in the first days of zebrafish development, for instance. However, the success of these approaches depends on the performance of the microscope.
|
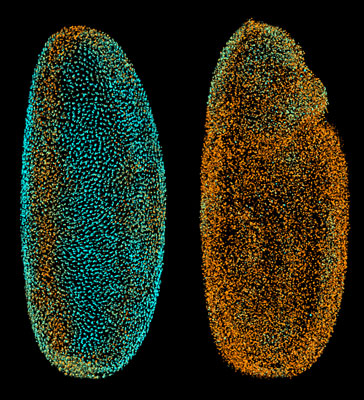 |
| The Fly Digital Embryo at different developmental stages, with cell nuclei coloured according to how fast they were moving (from blue for the slowest to orange for the fastest). The fruit fly embryo is magnified around 250 times.
|
|
'Non-transparent (i.e. opaque) samples like the fruit fly embryo scatter light, so the microscope picks up a mixture of in-focus and out-of-focus signals - good and bad information, if you like,' explained co-author Ernst Stelzer, a scientist at the European Molecular Biology Laboratory (EMBL) who took part in the research. 'Our new technique enables us to discriminate between that good and bad information.'
|
|
A couple of years ago, the scientists behind this new technique had already developed a novel microscope using fluorescence and thin laser beams. They pushed the method further to improve image quality and contrast by illuminating the specimen with a pattern of stripes instead of a homogeneous light sheet. The researchers captured multiple images of different phases of the light pattern; a computer combined these images, eliminating the effects of scattered light and generating a super-neat image of the sample.
|
|
The scientists combined this approach with imaging along different angles, and obtained two remarkable results. Firstly, they took around 1 million images to capture the first 3 days of zebrafish development from 3 different angles, generating films in which cell movements and the formation of the animal's eyes and midbrain are clearly visible. Secondly, even though the fruit fly embryo is opaque, they managed to produce spectacular three-dimensional (3D) time-lapse videos showing clearly the cellular dynamics of its development - all without damaging the specimen with excessive light exposure.
|
|
Not only are these images interesting to watch but they are also useful to other scientists. They can be used to run computational analyses following cell movements and divisions, and to create a scaffold for the mapping and comparison of gene expression in an organism.
|
|
Dr Stelzer underlined that the new method 'allows us to record organisms which have so far been poorly studied, because of their unfortunate optical properties.'
|
|
Led by researchers from the Heidelberg branch of the European Molecular Biology Laboratory (EMBL) in Germany, key contributions were made by researchers from Germany's University of Heidelberg, Karlsruhe Institute of Technology and Goethe University as well as from the Howard Hughes Medical Institute and the Sloan-Kettering Institute, both located in the US.
|
|
All data, images and videos are freely available online, alongside the data from the digital embryo, at www.digital-embryo.org.
|

