| Apr 11, 2011 |
New method for self-assembling molecules
|
|
(Nanowerk News) Researchers at the University of Sheffield have discovered a new way of making small molecules self-assemble into complex nanopatterns, which will push the limits of what is possible in `bottom-up´ methods of nanopatterning for advanced functional materials through molecular self-assembly.
|
|
The research, which was led by Dr Xiangbing Zeng and Professor Goran Ungar from the Department of Materials Science and Engineering, and colleagues from Martin Luther University Halle-Wittenberg in Germany, is published in Science (11 March 2011).
|
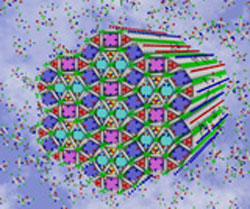 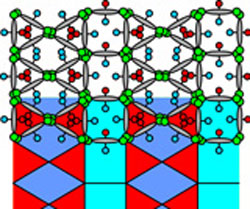 |
| Left: Artist impression of the complex multicolour honeycomb liquid crystal structure. Right: Another type of liquid crystal honeycomb formed by the same molecules but at a lower temperature.
|
|
The study opens the way to new methods of producing `bottom-up´ ultra-small electronic and photonic integrated circuits. This would mean that instead of the expensive and slow electron, ion-beam or X-ray lithography, the molecules would assemble and form the desired patterns themselves. Today visible or UV light is still used, but how small a pattern can be made is limited by the wavelength of light, that is of the order of a micron.
|
|
Solid-state materials, particularly those performing useful functions, often have their atoms and molecules arranged in rigid frameworks whose shapes are determined by the fixed lengths and angles of the strong chemical bonds that tolerate little change. Each solid-state framework structure is specific to the individual chemical compound. In contrast, in soft materials such as liquid crystals, the shapes of the frameworks are more easily adjustable. Self-assembling liquid crystals molecules usually contain two such incompatible parts that try to move away from each other, but cannot as they are chemically linked. As a result, such materials form a limited range of closed or open-cell structures.
|
|
This research demonstrates how increasing the number o carefully selected linked incompatible groups substantially broadens the range of cellular morphologies.
|
|
The discovery was made after the researchers used X-ray diffraction on very thin films (less than 1/10000 of a millimetre), using the beam of the Diamond Light Source synchrotron near Oxford. Other methods were also used to confirm the findings, such as neutron diffraction and atomic force microscopy, where a nanometre sized tip is used to gently tap over the sample surface "feeling" the individual molecules.
|
|
The nanopatterns presented by the team are the most complex patterns in liquid crystals so far. The findings show how the molecules have formed liquid crystal honeycomb shapes with highly complex tiling patterns with cells of up to five different compositions or `colours´ and five different polygonal shapes. The patterns have been formed by the molecules themselves finding the optimum stacking mode through countless trials and errors. However, a good understanding of the principles of self-assembly, gained by research such as this, allows chemicals to design molecules that would assemble precisely in the way they want.
|
|
|
|
The research also vividly illustrates the universality of physical principles, as it demonstrates how the same rules govern the structure, properties and phase transitions in such diverse systems as hard magnets and liquid crystals.
|
|
In magnets it is the magnetic moments of the electrons that flip, whereas in these liquid crystals the whole molecules turn around their axes. In the magnets the interactions are between magnetic fields, whereas in the liquid crystal it is the lack of miscibility between different molecular parts that is crucial, similar to the immiscibility between oil and water.
|
|
Professor Goran Ungar from the Department of Materials Science and Engineering at the University of Sheffield, said: "These findings are a new step toward making molecules that will create complex molecular electronic and photonic devices spontaneously and en masse.
|


