| Sep 22, 2014 |
The accelerator of molecular motors
|
|
(Nanowerk News) Peroxisomes are vital cell components that degrade cellular toxins and long-chain fatty acids. Their malfunction may result in severe, often lethal disorders. RUB researchers (Medical Faculty) have been studying the precise working mechanisms of peroxisomes for 25 years. In collaboration with Dortmund-based institutes at ISAS (Leibniz-Institut für Analytische Wissenschaften), they have successfully identified the "molecular accelerator" that activates the peroxisomal processes. To their surprise, it turned out to be an old acquaintance: a certain module of the familiar protein Pex22p, which has hitherto always been considered an anchor protein. The researchers report their findings in PLoS One ("The Cytosolic Domain of Pex22p Stimulates the Pex4p-Dependent Ubiquitination of the PTS1-Receptor").
|
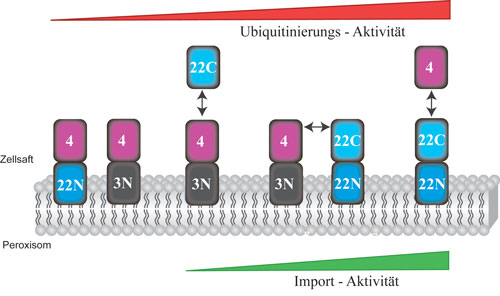 |
| Modular concept of peroxisome fusions: the soluble Pex22C component accelerates Ub-activity of the peroxisomal Ub-machinery even without the anchor component Pex22N, thus enabling the import of enzymes into the peroxisome.
|
|
Essential: importing enzymes into peroxisome
|
|
Peroxisomes are of vital importance for the enzymatic degradation of long-chain fatty acids and cellular toxins. In order for them to fulfil this function, the relevant enzymes have to be imported into the peroxisomes first. The bulk is brought into a peroxisome by the import receptor Pex5p. That receptor, in turn, is regulated by the protein ubiquitin (Ub): the modification of the receptor with an Ub-molecule paves the way for a new import reaction for further enzymes that have to be transported into the peroxisome.
|
|
Misjudged as anchor
|
|
In their previous projects, the team of Jun.-Prof. Harald Platta, Dr. Fouzi El Magraoui and Prof. Ralf Erdmann has already defined the basic composition of the peroxisomal Ub-machinery. They identified six related proteins and subdivided them into three functionally diverse groups. "It has remained unclear, however, how that molecular machinery is activated resp. how its activity is boosted," explains Harald Platta. In the current study, the researchers have succeeded in identifying the module that fulfils this vital function. "To our surprise, it's turned out to be not a hitherto unknown protein, but the component of a familiar protein, the already identified Pex22p," says Platta. "We have previously assumes that Pex22p's function is limited to being an anchor protein and that, by binding the soluble Pex4p to the membrane of the peroxisomes, it supports its function indirectly."
|
|
Research based on the modular concept
|
|
In order to ascertain which components are relevant, the researchers deployed the modular concept to assemble them by means of genetic fusions in various combinations. In doing so, they found out that the membrane anchor component of Pex22p was entirely irrelevant, provided that the soluble Pex4p was bound to the peroxisome at the membrane anchor component of Pex3p that was not functionally involved in the import reaction. This combination shows a low Ub-activity, which is too insignificant to modify a sufficient number of import receptors and to trigger the import of enzymes into the peroxisome. It isn't until Pex22(C) is added that Ub-activity increases, thus enabling the import of a sufficient number of enzymes into the peroxisome, in order to guarantee the peroxisome's functionality.
|
|
Not merely looking for new components, but also revising familiar ones
|
|
The discovery of this "accelerator" of the peroxisomal Ub machinery and – linked to it – the import machinery is relevant not just for understanding peroxisomal disorders such as the Zellweger syndrome. "It turns out that certain central proteins can fulfil several important tasks in biochemical systems in general," explains Harald Platta. "When analysing the molecular basis of various biochemically defined disorders, it will become relevant in general to identify a system's lacking activities not just by searching for new, unknown proteins. Our study demonstrates that a lacking function such as this may be perhaps already 'concealed’ in a familiar protein."
|

