| Nov 19, 2014 |
A better way to design genes
|
|
(Nanowerk News) A new web tool allows researchers to obtain higher protein yields for a range of life sciences applications including biosensing and drug manufacturing ("Codon Optimization OnLine (COOL): a web-based multi-objective optimization platform for synthetic gene design"). Designed by scientists at A*STAR, the program provides users with a simple and flexible interface for optimizing the sequence of synthetic genes for improved expression of desired proteins.
|
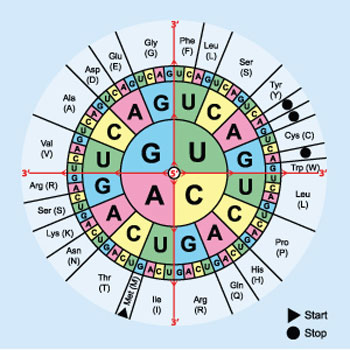 |
| Multiple DNA triplets can code for the same protein building block. The process of picking the best triplet for the host organism is known as codon optimization. (Image courtesy of Mouagip)
|
|
Through millions of years of natural selection, genes have evolved to allow for optimal protein expression. This includes a preference for certain DNA triplets, called codons, over others because of the relative abundance of the molecules involved in protein production. As a consequence, when biologists take genes from one organism and insert them into another to create synthetic genetic circuits, the gene sequences are often poorly adapted for the protein machinery of the new host. That is why scientists must go through a process known as codon optimization to tailor the DNA specifically to the new host organism. Most existing tools for codon optimization have fairly limited design criteria, however, and their rigid algorithms often fail to find the best sequence for a particular application.
|
|
Dong-Yup Lee, together with colleagues Ju Xin Chin and Bevan Kai-Sheng Chung from the A*STAR Bioprocessing Technology Institute in Singapore, sought a more adaptable solution for codon optimization. They designed ‘Codon Optimization OnLine’, or COOL, a user-friendly platform that incorporates customizable design parameters to find the optimal gene sequence. The tool also supports a wide range of visualization capabilities, which means researchers can graphically compare the quality of various optimized DNA sequences.
|
|
“COOL is the first web application that provides a full suite of optimization options as part of a multi-objective framework,” says Lee. “We developed novel computational algorithms for codon optimization and implemented it into COOL so that any scientists in the bioengineering, biotechnology and synthetic biology fields can easily use it for their gene design.” In his own lab, Lee says that he has designed several gene sequences using COOL and “obtained very good results.”
|
|
The program provides suggested codon usage and codon pair patterns that correspond to efficient expression for four hosts — these include two bacterial species (Escherichia coli and Lactococcus lactis) and two yeast species (Pichia pastoris and Saccharomyces cerevisiae) — all of which are commonly used in synthetic biology. Researchers can also input reference gene sets manually and Lee’s team is currently extending the built-in expression systems to include mammalian and plant cells.
|

