| Jul 16, 2015 |
Editing the genome with light
|
|
(Nanowerk News) A research group at the University of Tokyo has developed a new technology to optically control genome editing (Nature Biotechnology, "Photoactivatable CRISPR-Cas9 for optogenetic genome editing").
|
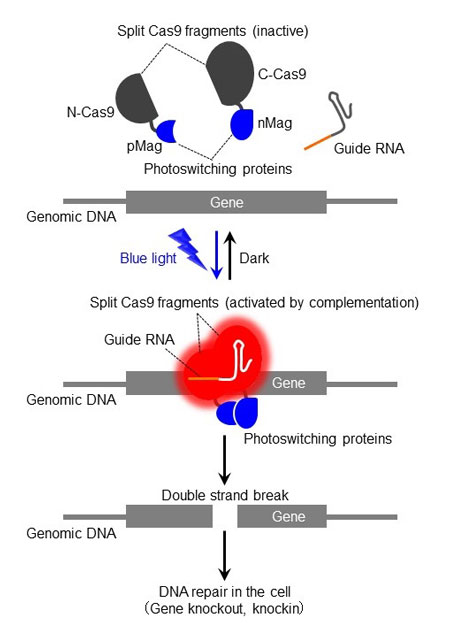 |
| Photoactivatable Cas9 (paCas9) system allows optical control of genome editing. N- and C-terminal fragments of Cas9 (N-Cas9 and C-Cas9) showing no significant catalytic activity were fused with photoswitching proteins (pMag and nMag). Upon irradiation with blue light, N-Cas9 and C-Cas9 bind to each other in response to light-dependent interaction of pMag and nMag. This allows the activation of the split Cas9 fragments, leading to site-specific cleavage of user-defined DNA sequences in the genome (switch-on). When the irradiation is turned off, N-Cas9 and C-Cas9 dissociate and lose catalytic activity (switch-off). (Image: Moritoshi Sato and Yuta Nihongaki)
|
|
To properly edit the genome, it is necessary to cut user-defined sites in the genome with a DNA nuclease. However, because the activity of conventional DNA nucleases cannot be controlled at all, it is not possible to cut the targeted DNA only in the desired cells and at the desired time. These limitations have constrained the possibilities of genome editing technology.
|
|
Ph.D. course student Yuta Nihongaki, Associate Professor Moritoshi Sato and their research group at the Graduate School of Arts and Sciences have developed a new technology to manipulate the activity of a DNA nuclease (Cas9) derived from the bacterial CRISPR-Cas9 system by using photoswitching proteins that can be controlled by blue light.
|
|
The new technology is expected to overcome the limitations of conventional genome editing technology and should contribute to expanding potential biomedical, pharmaceutical and agricultural applications including development of new cultivars.
|

