| Jun 19, 2017 |
Figuring out the 3-D shape of molecules with a push of a button
|
|
(Nanowerk News) An international team of researchers led by Carnegie Mellon University chemist Roberto R. Gil and Universidade Federal de Pernambuco chemist Armando Navarro-Vázquez has developed a program that automates the process of figuring out a molecule's three-dimensional structure. The technique, described in a paper in Angewandte Chemie ("Computer-Assisted 3D Structure Elucidation of Natural Products using Residual Dipolar Couplings"), compresses a process that usually takes days into minutes and could shorten the pipeline of drug discovery by reducing human error.
|
|
Figuring out the chemical structure of a molecule from scratch is an essential part of researching chemicals that come from nature, or "natural products." For substances with possible pharmaceutical use, that structure can reveal how the substance might interact with the human body.
|
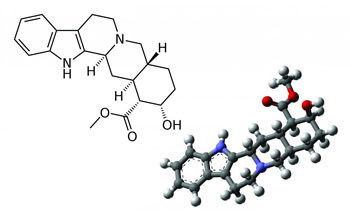 |
| A computer program can figure out the 3-D structure (right) of molecules like Yohimbine with the push of a button. Molecules with the same 2-D structure (left) can have different 3-D structures. (Image: 2D structure: acdx (Wikipedia) 3D structure: MindZiper (Wikipedia))
|
|
"If the molecule is going to be a drug, you need to know the shape of the molecule to know how it's going to interact with a receptor," says Gil, a professor in the Department of Chemistry at Carnegie Mellon's Mellon College of Science.
|
|
The first step in determining a molecule's structure is determining its atomic building blocks, followed by finding out its two-dimensional structure, which shows how each atom is connected to each other. While some atomic bonds are stiff, others can rotate around a joint, making it possible for molecules with the same components and two-dimensional (2D) structures to have different three-dimensional (3D) shapes.
|
|
Small differences in shape can translate into large changes in how drugs act in the body. For instance, rotating one bond in the popular pain reliever ibuprofen makes it completely inactive. Similarly, starch and cellulose share the same 2D structure but have different 3D shapes. That difference is the reason why humans can digest grains and not wood.
|
|
Gil and Navarro-Vázquez have been working for eight years to simplify the process of finding and sorting through the possible 3D shapes for any given 2D structure. And with the laboratory tools for gathering data on a molecule's 3D structure becoming more widespread and available, the time was right to develop a method to automate and streamline the process.
|
|
The researchers created a program, written in the Python programming language, that makes use of Residual Dipolar Coupling (RDC) information, a measure of the distance between atoms extending from rotating bonds. Fed with data about a given molecule from RDC experiments, the program generates possible ways that the molecule can exist in three dimensions, and picks the most likely option.
|
|
The technique is most effective at tackling the 3D structure of organic molecules that are small-to-medium sized, and relatively rigid, with carbon atoms packed into rings rather than linked into long, bendy chains. The team tested their program on six such molecules, including naltrexone, a medication used to block the effects of opioids, and strychnine, a pesticide.
|
|
First, they determined the 2D structure of each molecule using a computer-assisted structure elucidation (CASE) program, aided by a collaboration with Clemens Anklin, vice-president for NMR applications and training at Bruker Corporation. They fed that information, along with RDC data about the molecule, into their new program. In each case, their program was able to pick out the correct 3D structure.
|
|
"You push a button, and with little human or no human intervention, you go from 2D to 3D structure in one shot," says Gil.
|
|
But just as important as the program's speed is its thoroughness.
|
|
"The amount of natural products that are wrongly reported, where the reported structure does not correspond to the real structure, is really large," says Navarro-Vázquez, a chemistry professor at Universidade Federal de Pernambuco in Brazil. This program could be a valuable way to check for possible structures that researchers might otherwise miss, helping them avoid their own inherent biases.
|
|
Gil traces his vision of this process back to his time as a graduate student and a conversation with a mentor 30 years ago about the idea that chemists could one day put a substance into a machine and see its structure with the push of a button. Soon, he hopes, programs like his and Navarro-Vázquez's creation will integrate the earlier steps of the process, bringing that vision even closer to reality.
|
|
"We are very close," says Gil. "This is the dream of any chemist."
|

