| Jan 24, 2019 |
How bacteria build hyper-efficient photosynthesis machines
|
|
(Nanowerk News) Researchers facing a future with a larger population and more uncertain climate are looking for ways to improve crop yields, and they're looking to photosynthetic bacteria for engineering solutions.
|
|
In the Journal of Biological Chemistry ("The small RbcS-like domains of the β-carboxysome structural protein, CcmM, bind RubisCO at a site distinct from that binding the RbcS subunit"), a Canadian research team reports on how cyanobacteria finesse one of the most wasteful steps in photosynthesis.
|
|
The study investigated the assembly of carboxysomes in which the bacteria concentrate carbon dioxide, boosting the efficiency of a critical enzyme called RubisCO.
|
|
"Essentially everything we eat starts with RubisCO," said Matthew Kimber, a professor at the University of Guelph in Ontario, Canada, and senior author on the recent paper.
|
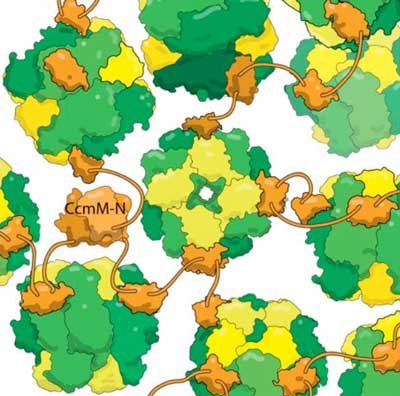 |
| New structural evidence shows that the cyanobacterial protein CcmM (orange) binds to fully assembled RuBisCo enzymes (yellow and green) without dislodging RbcS subunits (yellow). This crosslinked mixture forms the basis of the carboxysome, a compartment bacteria use to avoid photorespiration. Photorespiration is a wasteful step in photosynthesis caused when RuBisCo fixes oxygen instead of carbon dioxide, and is thought to reduce the yield of some crops by over 20%. By trapping carbon dioxide in carboxysomes, bacteria increase its concentration, helping RuBisCo correctly fix CO2. Greater insight into carboxysome assembly may help engineer higher-yield crops in the near future.
|
|
The enzyme, which is made of 16 protein subunits, is essential for photosynthesis. Using energy captured from light, it incorporates carbon dioxide into organic molecules from which the plant then builds new sugar. Unfortunately, it's not terribly efficient. Or, from Kimber's point of view, "RubisCO has a really thankless task."
|
|
The enzyme evolved in an ancient world where carbon dioxide was common and oxygen was rare. As a result, it isn't very picky in discriminating between the two gases. Now that the atmospheric tables have turned, RubisCO often accidentally captures oxygen, generating a useless compound that the plant then has to invest extra energy to recycle.
|
|
Compared with plants, cyanobacteria make very few such mistakes. This is because bacteria collect their RubisCO is into dense bodies known as carboxysomes. The bacteria pump bicarbonate (simply hydrated CO2) into the cell; once it gets into the carboxysome, enzymes convert the bicarbonate into carbon dioxide. Because the carbon dioxide can't escape through the protein shell surrounding the carboxysome, it builds up to high concentrations, helping RubisCO avoid costly mistakes.
|
|
Kimber's interest in carboxysomes is mainly in understanding the logic of their organization. "They're actually phenomenally intricate machines," he explains. "The cyanobacterium makes eleven or so normal-looking proteins, and these somehow organize themselves into this self-regulating mega-complex that can exceed the size of a small cell."
|
|
One of carboxysomes' most impressive tricks is self-assembly, which Kimber's lab set out to understand. They looked at a protein called CcmM, which corrals RubisCO enzymes into new carboxysomes. They knew that part of CcmM looks a lot like a subunit of RubisCO--so much so, in fact, that researchers suspect that ancient cyanobacteria originally created CcmM by duplicating a RubisCO gene.
|
|
Most scientists in the field believed that CcmM binds to the enzyme by usurping that subunit's spot. But when Kimber's lab took a detailed look at CcmM, using biophysics techniques to observe the protein's structure and binding, the results showed that received wisdom was wrong. True, CcmM was similar in shape to the small RubisCO subunit. But the complexes it formed still included all 8 small subunits, meaning that instead of stealing a spot from a RubisCO subunit, CcmM had to be binding somewhere else altogether.
|
|
"This is very odd from a biological perspective, because if CcmM arose by duplicating the small subunit, it almost certainly originally bound in the same way," Kimber said. "At some point, it must have evolved to prefer a new binding site."
|
|
The researchers also found that a linker between binding domains in CcmM is short enough that "instead of wrapping around RubisCO, it tethers (individual enzymes) together like beads on a string. With several such linkers binding each RubisCO at random, it crosslinks everything into this big glob; you wrap a shell around it, and this then becomes the carboxysome."
|
|
Scientists at a different university reported last fall that they had succeeded in making tobacco plants with a stripped-down carboxysome in their chloroplasts. Those plants didn't grow especially well, and the authors concluded that they had taken away too many components of the carboxysome; although it could be built in the chloroplast, it was a drag on the plants instead of a help.
|
|
Having a better understanding of how proteins like CcmM contribute to carboxysome construction and function could help bioengineers leverage carboxysome efficiency in the next generation of engineered plants.
|

