| May 08, 2013 |
Separation of dicarboxylic acids through molecular recognition and mechanochemistry
|
|
(Nanowerk News) How does one separate a mixture of components with very similar properties? In the journal Angewandte Chemie ("Dynamic Molecular Recognition in Solid State for Separating Mixtures of Isomeric Dicarboxylic Acids"), Croatian researchers have introduced a new approach to the separation of organic compounds. In their process, a “host compound” recognizes the desired “guest molecules”, not only in solution, but also when the host and mixtures of competitive guest are milled together in the solid state. For the separation of maleic acid, this recognition through mechanochemistry delivers selectivity equal to that achieved by crystallization from a solution.
|
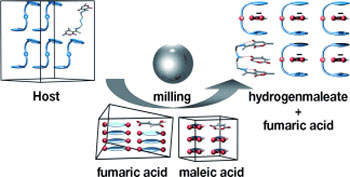 |
| Flexible even if constrained: A polyamine host recognizes dicarboxylic acids in solution and in solid state with the highest selectivity towards maleic acid, binding it from mixtures with up to five other dicarboxylic acids, including fumaric acid (see picture). Recognition using mechanochemistry is a dynamic process involving intermediate phases, resulting in the same selectivity as achieved by crystallization from solution.
|
|
Mixtures of dicarboxylic acids can be separated through crystallization, rectification, or extraction, processes often pushed to their limits because the dicarboxylic acids being separated have very similar chemical properties. Krunoslav Uzarevic and his co-workers at the Ruder Boskovic Institute and the University of Zagreb have now proposed an alternative method: separation through solid-state molecular recognition. Molecular recognition is the interaction of molecules that “fit” together without chemical bonding. Such interactions are known from phenomena such as enzyme recognition and DNA base pairing. This principle can also be used for the construction of supramolecular systems for nanotechnology. If one molecule is “lodged” inside another, the system is described as a host–guest interaction. Usually, molecular recognition is accomplished in solution, where the host and guest have the necessary mobility to come sufficiently close to one another and interact.
|
|
Separation of dicarboxylic acids by means of a host–guest interaction is challenging because the mixtures include isomers or molecules that are very geometrically similar. This is the case for two important dicarboxylic acids, fumaric acid and maleic acid. Fumaric acid is an intermediate product in the energy metabolism of all cells. It has many applications in the food and pharmaceutical industries. Maleic acid is used in polymer production, dying cotton, and decalcifying agents. Fumaric acid is usually obtained through the isomerization of maleic acid, which is harmful to health, so all residues of maleic acid must be scrupulously removed from the fumaric acid product.
|
|
As the host molecule for their separation, the research team selected a polyamine that consists of two oxygen-containing, six-membered, hydrocarbon rings bound together through a hydrocarbon chain that contains three amino groups. This compound is flexible and can crystallize in different conformations. This host molecule preferentially binds maleic acid. It grabs the maleic acid molecule between its two rings like a pair of tongs. It also selectively binds maleic acid in an excess of fumaric acid or four other related dicarboxylic acids. What is unusual is that this host–guest interaction works under conventional conditions, that is, when crystallizing the compounds out of a solution; as well as in the solid phase, when the dicarboxylic acid mixture is intensively milled together with the host molecule.
|
|
This type of selective binding and separation of guest molecules out of solid mixtures is, with the exception of a few pioneering experiments, a largely unexplored area of research. This work demonstrates that there is a vast potential of the solid-state molecular recognition for more environmentally friendly separation technologies.
|

