| Posted: Aug 15, 2014 | |
Direct and label-free discrimination of cytosine and its modifications utilizing silver ions in nanopores |
|
| (Nanowerk Spotlight) Cytosine (C) modifications such as 5-methylcytosine (mC) and 5-hydroxymethylcytosine (hmC) are important epigenetic markers associated with gene expression and tumorigenesis. However, bisulfite conversion – the gold standard methodology for mC mapping – can not distinguish mC and hmC bases. | |
| Recent studies have demonstrated hmC detection via peptide recognizing, enzymes, fluorescence and hmC-specific antibodies – nevertheless, a method for directly discriminating C, mC and hmC bases without labeling, modification and amplification is still missing. | |
| The alpha-hemolysin (α-HL) nanopore has been employed for DNA sequencing, various single-molecule detections and biomolecular interactions, by utilizing its narrowest constriction site. The latch zone (the vicinity of the vestibule constriction) of the α-HL pore was recently found to constitute a sensing region. | |
| This previously unrecognized sensing region was found to be able to detect individual abasic sites in dsDNA (see "Base-Excision Repair Activity of Uracil-DNA Glycosylase Monitored Using the Latch Zone of α-Hemolysin"). | |
| Inspired by this ideal, Dr. Wang and his collaborators have designed three dsDNAs containing C-C, mC-C or hmC-C at the 10th base-pair, which will place these mismatches at the latch zone of the nanopore for discrimination. | |
| "We found that the residual current follows C-C > mC-C > hmC-C, which suggest that the cytosine may block less ion current flow (higher residual current) at the latch zone compared to mC and hmC. This is because mC has an extra methyl group, while hmC have extra methyl group and then a hydroxy group, which could block more ion current," says Dr. Wang. | |
| The duration follows mC-C > C-C > hmC-C, which suggest that mC-C duplex is the most stable pair, then follows the C-C pair and the hmC-C is the least stable pair. | |
| These results, which we have reported in the August 8, 2014 online edition of Scientific Reports ("Single Molecule Investigation of Ag+ Interactions with Single Cytosine-, Methylcytosine- and Hydroxymethylcytosine-Cytosine Mismatches in a Nanopore") demonstrate that single base of C, mC and hmC can be discriminated at the latch zone of the nanopore. | |
 |
|
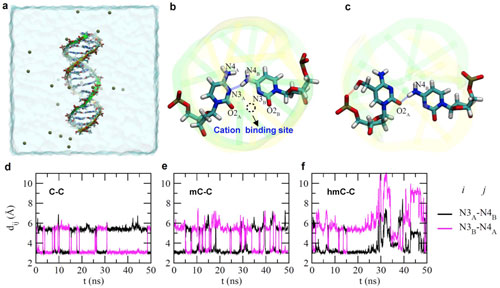
| Molecular dynamics simulations of DNA duplex containing C-C, mC-C and hmC-C mismatches. (a) (LBQ is the creator of figure 5a). Sideview of the simulation system. The DNA duplex is in the 'stick' presentation and two backbones are illustrated as yellow and green belts respectively. Potassium ions that neutralize the entire simulation system are shown as tan balls. Water in a cubic box (78.5 x 78.5 x 78.5 Å3) is shown transparently. (b) A snap-shot of pairing between two cytosine bases. The dashed circle highlights the binding site for a cation. (c) A snap-shot of hmC-C pairing before the pairing was broken. (d–f) Time-dependent distances between the N3 atom of one base and the N4 atom of the other base, in C-C (d), mC-C (e) and hmC-C (f) mismatches. (Reprinted with permission by Nature Publishing Group (click on image to enlarge) | |
| The interaction of DNA in the α-HL pore can also be modified by the specific interaction of Ag(I) (Ag+) with C-C mismatch, which strongly stabilizes DNA duplexes. | |
| In order to confirm our previous results, we tested the effects of silver ion on the C-C, mC-C and hmC-C containing duplexes. We found that with the addition of Ag(I), the residual current differences with the addition of Ag- are C-C > mC-C > hmC-C. The dwell time differences (ratios) with the addition of Ag+ are also C-C > mC-C > hmC-C. | |
| We therefore concluded that the C, mC and hmC bases also can be discriminated with the effects of silver ion. | |
| The nature of coordination of Ag+ with C-C mismatches is under debate. | |
| Our molecular dynamics simulations reveal that the paring, via a hydrogen bond, of a C-C mismatch results in a binding site for cations, such as K+ and Ag+. It is a dynamic coordination between N3A and O2B, or N3B and O2A for C-Ag-C interactions. | |
| These dynamic interactions suggest that the coordination of Ag+ in C-Ag-C complexes may have a different mechanism. | |
| Our result also is confirmed by melting temperature (Tm) measurements. | |
| With these two key differentiators (durations and residual currents differences), we can directly discriminate C, mC and hmC bases in the nanopore without amplification and modification. | |
| Thus our study demonstrates both the elucidation of the biophysics of C-Ag-C binding within the nanopore cavity as well as demonstrates an innovative new detection principle for distinguishing important epigenetic markers. | |
| This approach might be extended to study other cytosine modifications, like 5-formylcytosine (5fC) and other metallo-pair interactions. It is possible that a target fragment of the genomic sample can be obtained by a suite of restriction endonucleases, which can be purified and segregated for nanopore research. | |
| In conclusion, we have introduced a new direct detection method, supported by simulation, to discriminate between C, mC, and hmC. This work shows direct applicability in studying future targets and biophysical interactions, as well as analyzing epigenetic markers in target gene sequences. | |
|
By Dr. Yong Wang, Nanopore Single Molecule Sensing Group, Biological Engineering and Dalton Cardiovascular Research Center, University of Missouri-Columbia.
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
