| Posted: Sep 18, 2014 |
Researchers design an 'artificial nose' to detect DNA differentiation with single nucleotide resolution
|
|
(Nanowerk News) This is a method of identification of nucleic acids, with single nucleotide resolution, based on the generation pattern (bar code) inspired by our olfactory system. The differences in trans-membrane transport can be used to generate fluorescence patterns. That allows the differentiation of molecules as DNA or RNA by means of pattern generation and/or recognition protocols.
|
|
This work, published in Small ("Single-Nucleotide-Resolution DNA Differentiation by Pattern Generation in Lipid Bilayer Membranes"), has been fully developed at CIQUS by the PhD student Juan M. Priegue in collaboration with the postdoctoral researcher Javier Montenegro, under the direction of Professor Juan R. Granja.
|
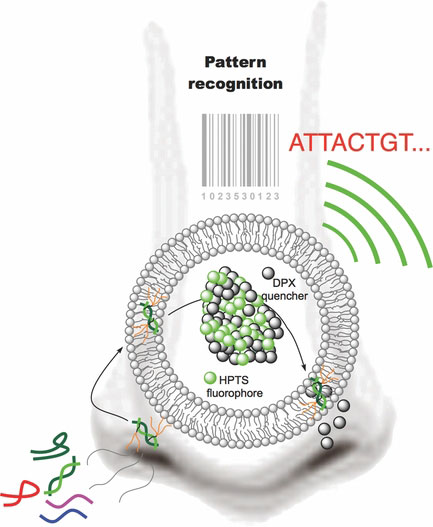 |
| Scheme: 'DNA nose' that identifies (smell) a variety of polynucleotides (Illustration: E. Vazquez, CIQUS)
|
|
Pattern recognition is the mechanism that operates in mammal’s olfactory reception. In effect, humans can detect thousands (or millions) of odorants with only hundreds of olfactory receptors. Incompatible with one to one recognition, the olfactory sensing system generates response patterns that configure a unique aroma sensation in the brain. In other words, in response to the detection of an odorant molecule the olfactory receptors “generate” a pattern that is recognized in the brain. This is called a pattern generation/recognition sensor.
|
|
Thus, this methodology has an enormous power for the amplification of the differences in very similar molecules. So it has inspired researchers to develop their own sensing system based for example in host/guest chemistry.
|
|
However, in this work researchers have employed the transport of DNA molecules across lipid membranes (as the cell membranes) to generate different transport patterns. They hypothesized that if a long DNA is transported fast and at low concentration, and a short DNA is transported slowly and at high concentration, we could use these different transport behaviours to detect and differentiate DNAs molecules.
|
|
The results of this study have shown that the amplification of differences observed during transport events allowed the differentiation of DNAs with an outstanding single nucleotide resolution. These findings are of interest because they might serve as a blueprint for the fabrication of many other pattern-based sensing systems in any other polymer of biological relevance (DNA, RNA...), a field that continues to be a major challenge in biochemistry.
|
|
Technical note
|
|
The detection and identification of polymers of biological interest -such as DNA or RNA- continues to be a major challenge in biochemistry. The transport of polyanions (i.e. DNA) across the membrane of fluorophore-loaded liposomes can be activated by small cationic amphiphilic molecules (activators). The corresponding fluorescent signals obtained in this type of transport experiments are different, depending on the molecules employed to activate DNA transport.
|
|
Therefore, these differences in transport can be used to generate fluorescence patterns that turn out to be distinctive for similar small non-ionic activators. Thus, this protocol allows the differentiation of similar analytes by means of pattern generation/recognition protocols.
|
|
In this new work researchers report that the same methodology can be also applied to generate unique dose-response patterns for different anionic polymers (DNA, RNA) of biological relevance. For that purpose, they have “de novo” designed and developed a synthetic strategy for the preparation of dynamic oximes-amphiphiles for the DNA transport activation across lipid bilayers. The fluorescent fingerprints (dose-response plots) generated in vesicle transport experiments allowed the differentiation of a collection of biopolymers with excellent reproducibility and precision, up to single nucleotide resolution of short single stranded oligonuclotides.
|

