| Posted: May 03, 2016 |
A new model for simulating DNA's 'atmosphere' of ions
(Nanowerk News) Nucleic acids, large biomolecules essential to life, include the familiar double-stranded deoxyribonucleic acid (DNA), a very stable long molecule that stores genetic information.
|
|
In nature, DNA exists within a solution rife with electrostatically charged atoms or molecules called ions. A recent study (Biophysical Journal, "The Role of Correlation and Solvation in Ion Interactions with B-DNA") by researchers at Pacific Northwest National Laboratory (PNNL) proposed a new model of how B-DNA, the form of DNA that predominates in cells, is influenced by the water-and-ions "atmosphere" around it.
|
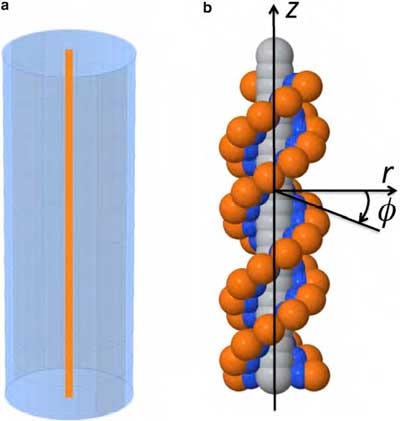 |
| The study compared two models of very large ions (macroions) that carry an electrical charge, and compared the results to experimental studies. On the left is a model of a cylinder with uniform axial charge density; on the right is the more complex (and useful) discrete charge model.
|
|
Understanding the ionic atmosphere around nucleic acids, and being able to simulate its dynamics, is important. After all, this atmosphere stabilizes DNA's structure; it impacts how DNA is folded and "packed" in cells, which triggers important biological functions; and it strongly influences how proteins and drugs bind to DNA.
|
|
The research combines theoretical modeling and experiments in a study of ion distribution around DNA. It was led by PNNL physical scientist Maria Sushko, computational biologist Dennis Thomas, and applied mathematician Nathan Baker, in concert with colleagues from Cornell University and Virginia Tech.
|
|
Earlier approaches have been used to simulate the distribution of ions around biomolecules like DNA. But only roughly. The PNNL-led study goes beyond commonplace electrostatics to propose a more refined but still computationally efficient model of what happens in these critical ionic atmospheres.
|
|
"The main idea was to dissect the complex interplay of interactions, and to understand the main forces driving ions deep inside the DNA helix and the forces keeping them on its surface," said Sushko, the paper's first author. That interplay includes the correlation of ions within the solution, how they move, how they interact with one another, and how they interact with the DNA.
|
|
The new model has two key advantages over older simulations: It allows researchers to turn ion-water and ion-ion interactions on and off at will. "We can calculate important interactions independently," she said, a flexibility not present in previous simulations. And the new model is computationally efficient, allowing researchers to cheaply simulate a large-scale molecular event over a long time scale.
|
|
Results
|
|
Importantly, both previous and new experiments by the Cornell colleagues identified the number of bound ions around DNA. Previous simplified models were also able to reproduce this number. But the new model "is richer than that," said Sushko, because it gives more details on how ions are distributed along the surface of DNA and within DNA's critical grooves. "DNA interaction will strongly depend on where those ions sit," she said. For one, the presence of ions in the grooves relates to how compact DNA will be. "The more ions within the grooves," said Sushko, "the more compact the structure."
|
|
The researchers confirmed that biological "correlation," a measure of ion affinity, allowed DNA to pack more tightly by effectively neutralizing DNA's electrostatic charge. Researchers also observed how ions get distributed through a solution, a water-ion interaction called solvation. The stronger the water-ion interaction, the larger the effective ion size, and therefore the less likely the ion was to settle in the DNA's grooves. More strongly solvated ions, therefore, create a different environment for DNA folding.
|
|
Researchers observed results regarding the activity of three types of salts within the simulated ionic environment. Small, single-charge ions did not strongly react with water; about 50 percent of these bond ions could penetrate into DNA grooves. Large ions with triple charges were not strongly hydrated, but their size prevented penetration into the grooves. ("They just decorate the surface," said Sushko.)
|
|
Only 15 to 20 percent of ions with double charges, which were strongly hydrated and strongly correlated, settled in DNA grooves. That showed a "very delicate interplay" of ion-to-ion and ion-to-water interactions, according to Sushko.
|
|
Why It Matters
|
|
These results highlight important aspects of the properties of electrolyte solutions influencing the ionic atmosphere that impacts DNA condensation. This "packing" of DNA, which is otherwise one of the longest molecules in nature, is essential to DNA's role in gene regulation. DNA condensation is also the key to protein binding and drug binding. It therefore points to practical applications in medicine and biotechnology.
|
|
This research also highlights the impact of the ionic atmosphere on the interaction between biomolecules and a ligand: that is, the molecule, ion, or protein that binds with a protein or the DNA double helix for some biological purpose.
|
|
But it is the "methodology itself," not the designed simulations of DNA, that is most important, said Sushko, in part because it provides a new computational model of how to see into complex molecular systems. "We get a better fundamental understanding of the important forces."
|
|
Methods
|
|
Researchers employed two coarse-grained models to simulate the DNA macroion, which is a large colloidal ion carrying a charge. The goal was to capture two versions of detail on how ions spread out in a solvent and how they interact with simulations of DNA topology.
|
|
One DNA model posited an infinitely long cylinder with a uniform charge density along one axis. Sushko called it "a very crude model used a lot in the past. It explains quite a lot about ion interactions, but it is deficient in some ways." The second, more complex "discrete charge" model posited three types of spheres in a helical array that mimics B-form DNA. It had a 3D-like character that allowed ions to penetrate into DNA grooves.
|
|
The DNA simulations were run through four computational models of classical density function theory to assess the energetics of different ion-DNA interactions. Results were also compared to data from what Sushko called "state-of-the-art experiments" that used anomalous small-angle x-ray scattering. This technique, used to investigate the spatial dimensions of structures in the nanometer range, always yields a lot of detail about how ions are distributed around a biomolecule.
|
|
The uniformly charged cylinder model was not good at simulating the ionic atmosphere around DNA. "This model is a very common simplification," said Sushko. "You get the same number of ions attached to DNA, but the distribution is completely wrong. In this model, ions will just sit somewhere on the surface."
|
|
But their more complex discrete charge model provided a much more naturalistic portrait of ion distribution in an ionic atmosphere. Its simulations showed ions both clinging to the helical DNA surface and also penetrating into the DNA's grooves. "The small details of ion penetration are very important for the way DNA will package the chromosome," she said.
|
|
What's Next?
|
|
Researchers plan to study the role of the ionic atmosphere in mediating interactions between DNA molecules. They also plan to extend their DNA model to include DNA sequence-specific effects, which often influence ion binding, and DNA sequence-dependent structural variations.
|

