| Oct 01, 2022 |
A nano-IR probe for proteins in liquid environments
(Nanowerk News) A new technique using infrared (IR) light from the Advanced Light Source (ALS) revealed how the self-assembly of proteins is affected by environmental conditions in a surrounding liquid (PNAS, "In-vitro investigation of protein assembly by combined microscopy and infrared spectroscopy at the nanometer scale").
|
|
This nanoscale probe of soft matter in a liquid matrix will facilitate advances in biology, plastics processing, and energy-relevant applications such as electrocatalysts and batteries.
|
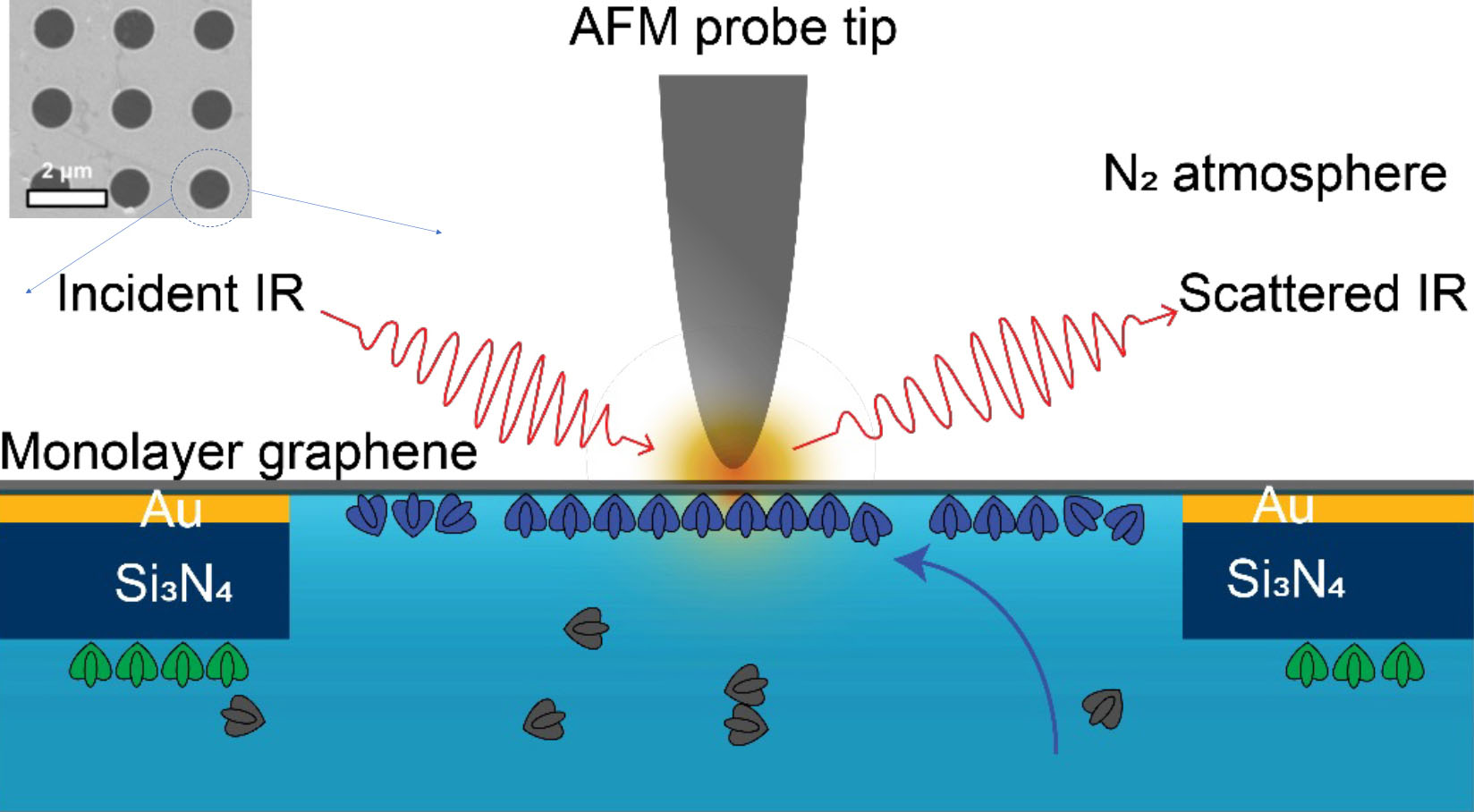 |
| Experiment schematic. The tip of an atomic force microscope (AFM) uses IR light, plasmonically enhanced at the apex, to probe the sample material in its liquid environment through a monolayer graphene window. The graphene is placed over 1 µm holes in a membrane of gold (Au) and silicon nitride (Si3N4). Only one hole is shown in the schematic; see inset for overview. (Image: Advanced Light Source)
|
More information, less damage
|
|
Proteins are essential to all living systems. They are also used in technological applications, including sensors, biocatalysts, and biomaterial synthesis. Most proteins organize into well-defined hierarchical structures, governed by dynamic noncovalent interactions (e.g., hydrogen bonding) that are sensitive to environmental conditions. This complexity calls for in vitro characterization tools that are noninvasive and provide nanoscale resolution.
|
|
Substantial effort has been put into developing microscopy tools to characterize protein assemblies, including x-ray crystallography, atomic force microscopy (AFM), fluorescence microscopy, and cryo-electron microscopy. However, these powerful imaging techniques can potentially damage or perturb delicate biological material and do not provide chemical information, preventing a fundamental understanding of the dynamic processes underlying their evolution under physiological conditions.
|
|
In this work, researchers addressed this challenge by developing a liquid cell enclosed by single-layer graphene, which serves as both a substrate and as infrared-transparent window.
|
Liquid-cell demonstration
|
|
To demonstrate the technique, the researchers used a combination of AFM topographic imaging at Berkeley Lab’s Molecular Foundry and synchrotron infrared nanospectroscopy (SINS) at ALS Beamlines 2.4 and 5.4. For the liquid cell, an atomically thin graphene layer was placed over a perforated layer of silicon nitride and gold. This separated the protein-buffer solution from the nitrogen-gas environment of the AFM tip. IR spectra (sensitive to molecular vibrational modes) were taken through the graphene at the perforations (1 µm in diameter).
|
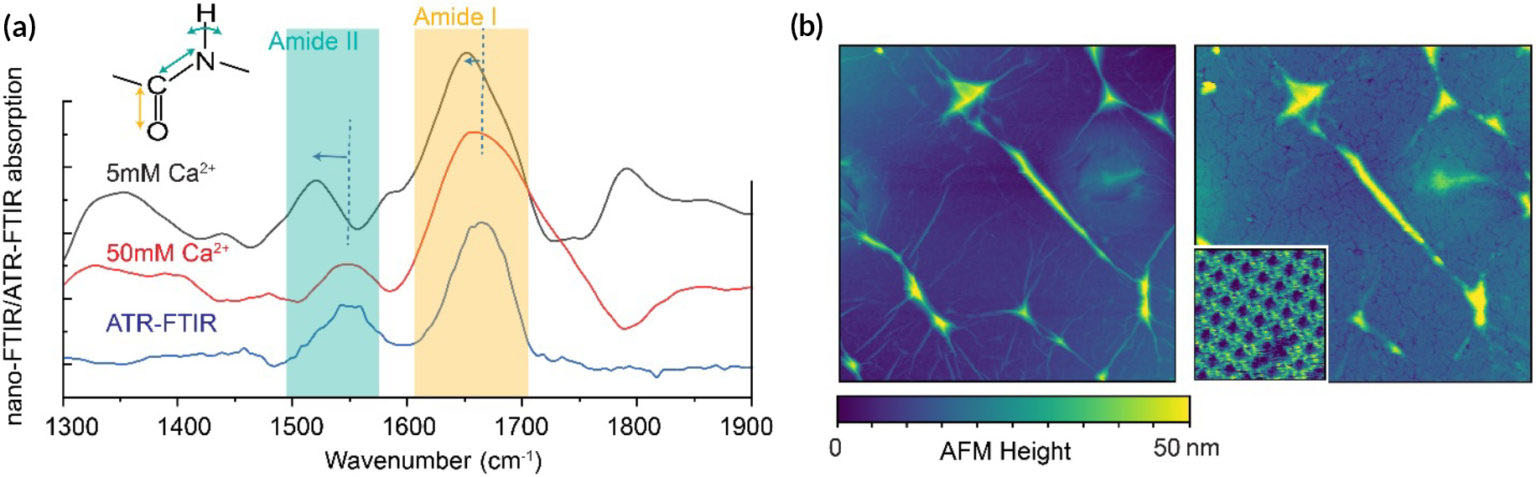 |
| (a) IR spectra of SbpA assemblies with various calcium-ion concentrations. Inset shows stretching and bending amide I and II modes. The ATR-FTIR curve (corresponding to dried SbpA assembled on graphene-coated gold film) serves as a reference. Relative shifts in peak positions are associated with hydrogen bonding, and peak-height ratios relate to bond orientations. (b) AFM topographic images of graphene membrane before (left) and after (right) SbpA assembly. Inset: Expanded image showing square lattice structure of the assembled proteins. Scan size: 3 µm for large images and 90 nm for inset. Bright lines are due to wrinkles in the graphene. (Image: Advanced Light Source)
|
|
The model system under study consisted of proteins (SbpA) that, in the presence of calcium ions, self-assemble into two-dimensional surface lattices. In many bacteria and archaea, these surface protein lattices serve as exoskeletons and sieves for nutrient uptake. The researchers were able to track the chemical and structural evolution of SbPA during self-assembly, with a resolution of 10 to 20 nm (on the order of the AFM tip size).
|
|
Different stretching and bending modes (the amide I and II bands) were recorded under different buffer conditions. The spectral intensities grew nonlinearly over time, matching the increase in ordered protein domains indicated by AFM images. After assembly, the spatial distribution of the amide band intensities revealed inhomogeneities among protein domains not visible in previously reported microscopy results. The work also showed that the evolution of the protein substructure depends on many environmental factors, including ionic strength, solvent, and substrate.
|
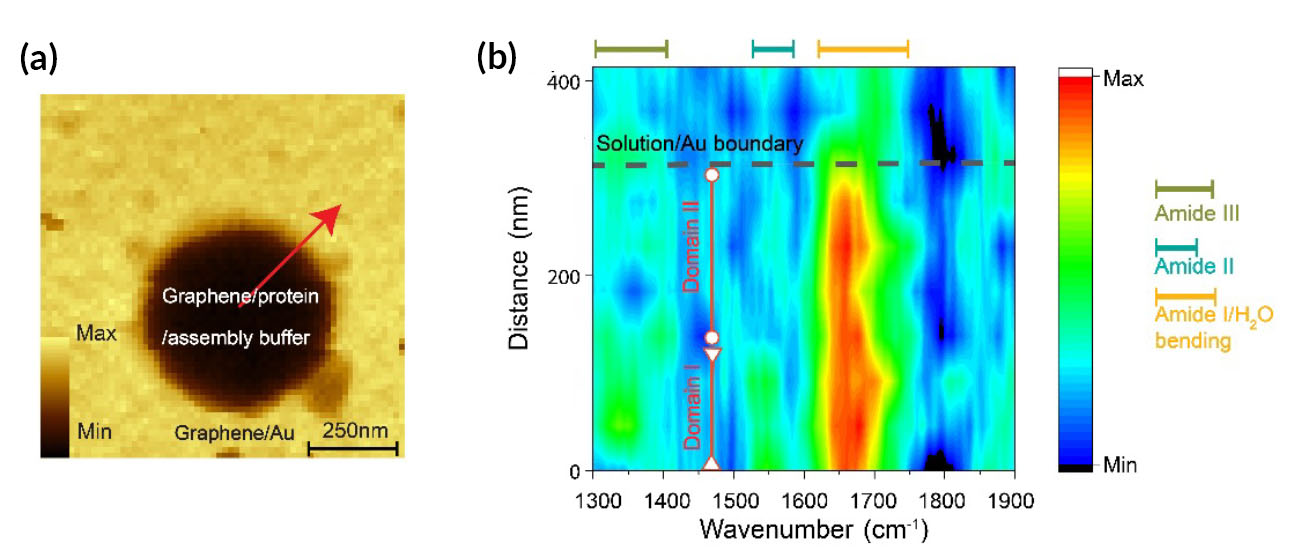 |
| Spatially resolved chemical mapping of SbpA protein assembly. (a) Image of the suspended graphene and surrounding area for a cell filled with SbpA in 50 mM Ca2+ solution. Total optical amplitude increases substantially when the probe is over the gold substrate. (b) Color map of regions of spectral intensity marking the amide I and II modes (as well as a third, low-frequency, amide III mode) along the red arrow in (a). Dotted black line indicates hole boundary; vertical red lines indicate different self-assembled protein domains. (Image: Advanced Light Source)
|
Bright future prospects
|
|
The combination of the graphene liquid cell and SINS offers enormous opportunities for operando characterization of proteins and other biological material. The researchers anticipate that this platform will also provide new molecular-level knowledge of many complex biological systems and processes, including the misfolding of beta amyloid proteins and the voltage-induced evolution of ion-channel proteins.
|
|
This new methodology also enables nondestructive spectroscopic imaging of other soft materials with nanometer spatial resolution, with examples ranging from biomaterial (enzymes and viruses) to polymer material, under in vitro conditions and external stimuli.
|
|
With advances in fabrication technology, the spatial resolution—correlated to the tip apex radius—can be further increased. The researchers also believe that dynamic infrared spectroscopic study of a single protein or DNA molecule during the execution of its biological function will be possible in the future.
|



