| Posted: Jul 15, 2017 | |
Sensing protein folding with nanopores(Nanowerk News) In a recent paper in ACS Nano ("Nanopore Sensing of Protein Folding"), researchers report results of all-atom molecular dynamics (MD) simulations that directly evaluated the effect of protein folding on the nanopore ionic current. |
|
| The team theoretically showns that measurements of nanopore ionic current can be used to monitor, in real time, folding-unfolding transitions of a protein and to characterize intermediate states along a protein folding pathway. | |
| The experimental detection of protein folding can be realized by collecting ionic current signatures of many proteins transiently passing, one-by-one, through the nanopore volume. | |
 |
|
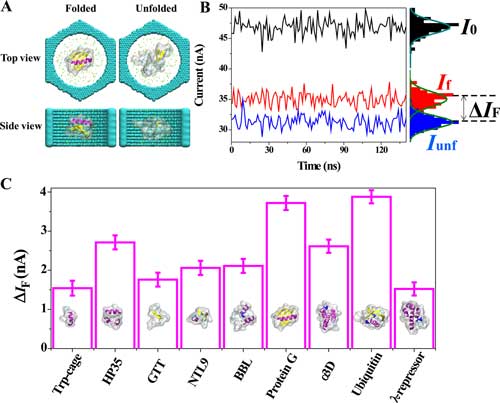
| Ionic current blockades report on the folding state of a protein. (A) A typical simulation system containing a folded (left) or unfolded (right) protein (protein G) at the center of a 6 nm-diameter solid-state nanochannel (cyan). An electric field is applied along the axis of the channel, and the protein is restrained to maintain its initial conformation. The protein is colored according to the secondary structure of its folded conformation: purple indicates α-helix, yellow indicates β-sheet, white and cyan indicate coil and turn, respectively. The semitransparent molecular surface represents the excluded volume of the protein. Green and yellow spheres depict potassium and chloride ions, respectively. Water is not shown for clarity. (B) Ionic current recorded from MD simulations of protein G placed in the nanopore in the folded (red) and unfolded (blue) states. The black trace indicates ionic current recorded in the absence of the protein. All-point histograms of the current values are shown at the right axis. The current traces were sampled every 2.4 ps and averaged in 1.2 ns blocks. The ionic current difference between the folded and unfolded states ΔIF = If - Iunf, where If and Iunf are the average currents in the folded and unfolded states, respectively. Data shown were obtained under applied electric field of 74.67 mV/nm, and the concentration of the KCl electrolyte was 2 M. (C) Blockade current difference (ΔIF) for nine proteins. Inset images illustrate the folded conformations of the proteins. All other conditions as in panel B. The eight additional simulation systems are shown in SI Figure S1. Error bars represent the standard error of ΔIF computed from the 150 ns ionic current traces averaged in 1.2 ns blocks. (© ACS) (click on image to enlarge) | |
| While offering high throughput, this approach, however, relies on statistical averaging over many protein translocation events, which eliminates some of the advantages of a single-molecule measurement. | |
| A much more attractive possibility, as the authors point out, is continuous monitoring of the ionic current through a nanopore that has a protein attached to its surface. Lipid-coated nanopore systems are particularly suitable for such measurements, as they can retain proteins within the nanopore volume long enough to detect a folding-unfolding transition in real time while allowing different copies of the proteins to enter and leave the nanopore. | |
| Decorated with metallic nanoantennas, plasmonic nanopores can be used to both rapidly change the temperature of a nanopore volume and to trap proteins, offering a single-molecule realization of a temperature jump experiment. | |
| By performing such ionic current measurements using nanopores of different sizes, one may be able to elucidate the effect of confinement on the protein folding process. | |
| "All of the above suggest that real-time monitoring of the protein folding-unfolding transition using an experimental nanopore system can be realized in the near future, leading to exciting developments in the area of molecular diagnostics and drug design," the scientists conclude their report. |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Subscribe to a free copy of one of our daily Nanowerk Newsletter Email Digests with a compilation of all of the day's news. |
