| Posted: Oct 26, 2017 | |
Engineering cell-membrane function with DNA origami nanodevices(Nanowerk News) Structural DNA nanotechnology, specifically the molecular self-assembly process known as DNA origami, has emerged as a versatile approach to fabricate nanodevices with complex nanoscale geometry, defined placement of molecular functionalities, and programed mechanical and dynamic properties. |
|
| Although prior work has utilized DNA strands bound to cell membranes to mediate target sensing or cell–cell binding, implementing DNA origami structures at the cell surfaces provides a distinct advantage for multiplexing function. | |
| In new work, reported in Advanced Materials ("Engineering Cell Surface Function with DNA Origami"), researchers from The Ohio State University establish a foundation for implementing the diverse function of DNA origami nanotechnology on cell surfaces by using hydrophobic anchors to attach 3D DNA origami nanostructures to the surface of five distinct cell types, including adherent, suspension, and primary cells. | |
| The team's method uses cholesterol-conjugated oligonucleotides as amphiphilic anchors that incorporate into the plasma membrane to facilitate installation of DNA origami structures onto the surface of living cells. This method is specific, reproducible, and reversible. | |
| "We demonstrate installation of a DNA nanoplatform that functions as a molecular-scale membrane-bound breadboard (MBB) that can be programed with multiple functions such as sequence-specific binding or fluorophores at defined locations," the authors write. | |
 |
|
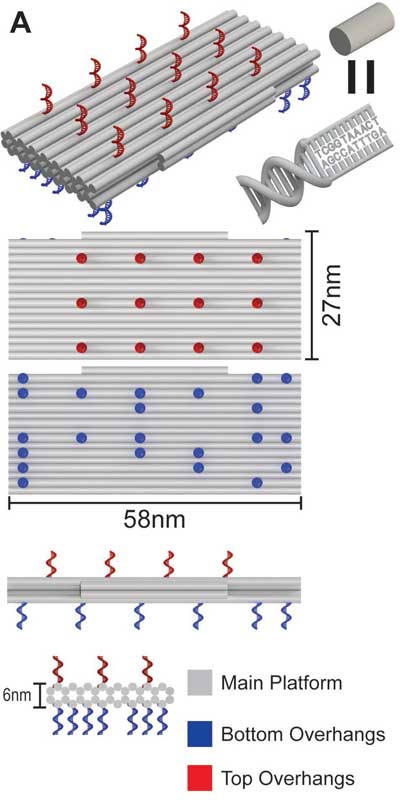
| Design and characterization of a molecular-scale membrane-bound breadboard. A) Isometric, top, bottom, and side views illustrate the structure design and locations of the top (red) and bottom (blue) overhangs. Gray cylinders represent dsDNA helices. (© Wiley VCH-Verlag) | |
| The sequence-specific binding allows the MBB to be used as a docking site for controlled spatial positioning of additional DNA-based constructs, devices, or higher order assemblies on the cell surface. | |
| The researchers also demonstrate this capability by programing both homotypic and heterotypic cell–cell binding where the sequence-specific DNA base pairing between MBBs on adjacent cells serves as a cellular Velcro (or “CellCro”). | |
| The characteristics of the MBB presented here, including the size and overhang binding specificity, could also allow for localization of many measurement functionalities on the same or distinct platforms on individual cells or imparting specific functions to certain cell types within mixed cell populations in lab-on-a-chip environments. | |
| More broadly, this work establishes a foundation to use DNA origami nanodevices as membrane engineering tools that can mimic and direct complex biological processes on the surface of various cell types in microtissue scale. | |
| "We envision that multifunctional DNA origami devices can transform the cell membrane into a programmable material that exploits the broad scope of biomolecular nanotechnology," the authors conclude their report. |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Subscribe to a free copy of one of our daily Nanowerk Newsletter Email Digests with a compilation of all of the day's news. |
