| Posted: Jul 02, 2007 | |
Developing nanotechnology noses to 'smell' cancer |
|
| (Nanowerk Spotlight) Biomarkers are of increasing importance in modern medicine for the purpose of early detection and diagnosis of a disease, for instance cancer. Biomarkers are mostly protein molecules that can be measured in blood, other body fluids, and tissues to assess the presence or state of a disease. For example, the presence of an antibody may indicate an infection or an antigen, such as PSA, might indicate the presence of prostate-specific cancer cells. Although protein-based approaches to early detection and diagnosis of cancer have a clear advantage over other, more invasive, methods, protein detection is a challenging problem owing to the structural diversity and complexity of the target analytes. State-of-the-art detection methods have limited application due to their high production cost and instability. Another limitation of current proteomic diagnostics is the limitation of arrays to one or a few markers only; in other words, you can only test for the specific markers that you are looking for and not generally measure levels of proteins in your blood in order to detect anomalies. A novel nanotechnology based protein detector array could change that. | |
| "The key finding of our work is that nanoparticles can be used to create selective receptors for proteins" Dr. Vincent M. Rotello tells Nanowerk. "Creation of an array of these receptors then provides a system where proteins can be identified, i.e. a chemical nose." | |
| Rotello is a Professor in the Department of Chemistry at the University of Massachusetts. Together with members from his group and the School of Chemistry and Biochemistry at the Georgia Institute of Technology, led by Professor Uwe Bunz, he just published an article in Nature Nanotechnology ("Detection and identification of proteins using nanoparticle–fluorescent polymer ‘chemical nose’ sensors") that describes how assemblies of gold nanoparticles with fluorescent PPE polymer provide efficient sensors of proteins, achieving both the detection and identification of analytes. | |
| "To my knowledge, this is the first use of the use of nanoparticles in array-based sensing" says Rotello. "We were motivated by the need for a general method for sensing and identifying proteins. While array-based sensing of proteins has been done before, our system is much more sensitive, and much more effective, i.e. we use fewer receptors to identify more proteins." | |
| There are two key challenge for the development of effective protein sensors: 1) the creation of materials featuring appropriate surface areas for binding protein exteriors, coupled with the control of structure and functionality required for selectivity. 2) The signal transduction of the binding event. | |
| "Nanoparticles provide versatile scaffolds for targeting biomacromolecules that have sizes commensurate with proteins, a challenging prospect with small molecule-based systems." explains Rotello. "Our strategy for the creation of protein sensors is to use the nanoparticle surface for protein recognition, with displacement of a fluorophore generating the output." | |
| To create the protein sensors, the scientists used six gold nanoparticle–fluorescent polymer conjugates. These particles serve as both selective recognition elements as well as quenchers for the polymer. The subsequent binding of protein analytes displaces the dyes, regenerating the fluorescence. | |
 |
|
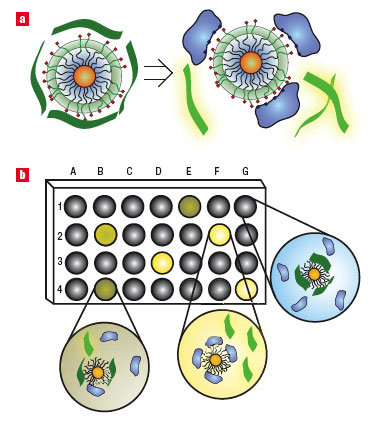
| Fluorophore displacement protein sensor array. a, Displacement of quenched fluorescent polymer (dark green strips, fluorescence off; light green strips, fluorescence on) by protein analyte (in blue) with concomitant restoration of fluorescence. The particle monolayers feature a hydrophobic core for stability, an oligo(ethylene glycol) layer for biocompatibility, and surface charged residues for interaction with proteins. b, Fluorescence pattern generation through differential release of fluorescent polymers from gold nanoparticles. The wells on the microplate contain different nanoparticle–polymer conjugates, and the addition of protein analytes produces a fingerprint for a given protein. (Reprinted with permission from the Nature Publishing Group) | |
| By modulating the nanoparticle–protein and/or nanoparticle–dye association, distinct signal response patterns can then be used to differentiate the proteins. The fluorescent indicator displacement assay does not require special instruments, and its sensitivity (due in large part to the high surface area provided by the nanoparticles) and speed facilitate protein detection. | |
| Rotello points out that real-world applications require identification of proteins at varying concentrations and of unknown identity. "Varying protein concentrations would be expected to lead to the drastic alteration of fluorescence response patterns for the proteins, making identification of proteins with both unknown identity and concentration challenging. To enable the detection of unknown proteins, we have designed a protocol combining LDA and ultraviolet (UV) measurements." | |
| In their experiments, the researchers also used a series of unknown protein solutions for quantitative detection. The unknown protein solutions were submitted to the testing procedures: of the 52 unknown protein samples, only three samples were incorrectly identified, affording an identification accuracy of 94.2%. | |
| "Our overall goal is to develop a more holistic approach to detecting diseases" says Rotello. "The plan is to monitor levels of proteins in the body, as opposed to looking for just a single marker." Of course, the key question is whether disease can effectively be 'smelled' – are there detectable differences in the relative ratios of proteins that can be used for diagnostic purposes?" | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
