| Posted: Feb 13, 2006 | |
Single Molecule Nanometronome |
|
| (Nanowerk News) Researchers at the Center for Biophysics and Computational Biology and Department of Physics at the University of Illinois at Urbana-Champaign and Howard Hughes Medical Institute constructed a DNA-based nanomechanical device called the "nanometronome". | |
| The device is made by introducing complementary singlestranded overhangs at the two arms of the DNA four-way junction. The ticking rates of this stochastic metronome depend on ion concentrations and can be changed by a set of DNA-based switches to deactivate/reactivate the sticky end. | |
| Reporting their work: "Single Molecule Nanometronome" in the Feb. 10, 2006 online edition of Nano Letters, Taekjip Ha and his colleagues describe that the device displays clearly distinguishable responses even with a single base pair difference. This may lead to a single molecule sensor of minute sequence differences of a target DNA. | |
| As Ha told Nanowerk, "As we watched our DNA device go "tick, tick, tick", we wondered if we could change how often the device ticks. This is precisely what we reported in this work, using additional DNA switching elements. We named it "single molecule nanometrome" because the ticking rate of this ten nanometers sized device is tunable, and the device was observed one molecule at a time." | |
 |
|
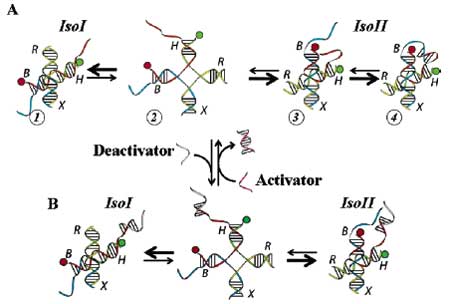
| (A) Diagram of conformational changes of a nanometronome. In terms of native four-way Holliday junctions, structure 1 possesses conformation IsoI whereas structure 3 and structure 4 possess conformation IsoII. Structure 2 is believed to be a transition or intermediate state during transitions. Base pairing of the sticky ends lowers the free energy of the structure 4 forcing the nanometronome to stay in conformation IsoII longer. The presence of Mg2+ is critical for the transitions to occur, and the rates of transition depend on [Mg2+]. (B) Reversibly switching sticky ends on and off by utilizing the short single-stranded deactivator/activator. The deactivator competitively binds onto the single-stranded overhang at the end of helix H, silencing the sticky ends. This binding leaves an overhang handle for the activator to bind and later remove the deactivator via three-stranded branch migration. (Courtesy of Taekjip Ha, University of Illinois at Urbana-Champaign.) | |
| According to the researchers, this is the first demonstration of a DNA-based nanodevice observed at the single molecule level. They learned that even a single base pair difference in the sticky end can be easily detected and single molecular heterogeneity is small enough that even at the single nanometronome level it should be possible to distinguish one base pair difference. Possible applications in the distant future may include the ultrasensitive detection of single nucleotide polymorphism. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
