| Posted: Aug 07, 2014 | |
Detecting damaged DNA with solid-state nanopores |
|
| (Nanowerk Spotlight) DNA is constantly being damaged in our cells by radiation and other random sources. There is a long list of structural and chemical variations to DNA that have impacts on human health. One of the major forms of this damage is called depurination, or the selective loss of A and G bases from the double helix structure. | |
| In our cells, there is a system in place to fix depurination, called the base excision repair (BER) pathway. It usually is quite successful at repairing the damage, but can sometimes make mistakes that result in mutations. As a result, depurination is directly linked to a host of diseases, including anemia and cancer. | |
| Better tools for the study of depurination could therefore be important in studying these diseases or potentially in their early diagnosis. | |
| In new work coming out of the Hall Lab at Wake Forest University School of Medicine, the group shows that DNA depurination can be detected electrically using solid-state nanopores. They reported their findings in PLoS ONE("Detecting DNA Depurination with Solid-State Nanopores"). | |
| Depurination has conventionally been studied indirectly – e.g. determining the replicative ability of DNA – or with expensive bulk techniques like liquid chromatography. | |
| "Our new technique is fast, requires little starting material, and can offer a rough assessment of depurination at the single-molecule level," Adam R. Hall, an Assistant Professor of Biomedical Engineering at Virginia Tech-Wake Forest School of Biomedical Engineering and Sciences, tells Nanowerk. "As with other single-molecule approaches, this allows the measurement of small populations that would be masked or averaged out in bulk techniques." | |
 |
|
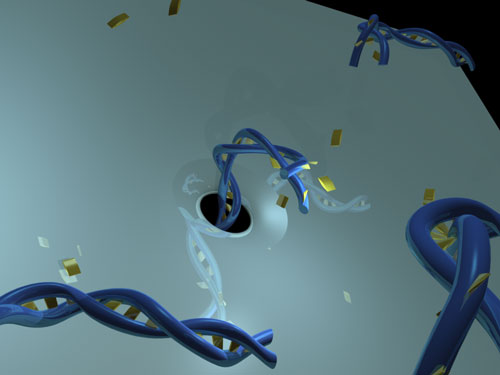
| Schematic of the system: a short duplex DNA strand is pushed through a nanopore. (Image: The Hall Lab, Wake Forest University School of Medicine) | |
| The team's solid-state nanopore devices are single fabricated, nanometer-scale (5-6 nm) holes in thin SiN membranes through which individual molecules can be threaded electrophoretically. As they pass, their presence and certain aspects of their structure can be determined, much like a film strip being run through a projector. | |
| "What we have found is that the structural changes resulting from depurination tend to act as 'breaks', slowing the threading process down," explains Hall. "The more damage is present in a molecule, the slower it moves through the nanopore, and this can be observed from the electrical signal." | |
| In their experiments, the researchers found that depurinated DNA translocates up to an order of magnitude more slowly than undamaged molecules, on average. | |
| These results advance the study of depurination and DNA damage by providing a new diagnostic tool that has extremely high sensitivity, is fast, label-free, and can be used for a coarse determination of either the overall level of depurination within a collection of dsDNA or the degradation of individual translocating molecules. | |
| As depurination is also a natural process, occurring at a certain rate over time, the scientists expect that his techniques might be useful in diverse areas like forensics. | |
| Hall points out that a great deal of focus has been put on solid-state nanopores as a potential way to sequence DNA ultrafast. While biological nanopores have made tremendous strides in this context, artificial nanopores have not been developed as rapidly. | |
| "This is due largely to limitations in fabrication and physical limitations of the system," he says. "However, we feel that there are plenty of areas that solid-state nanopores can access more readily- many with clinical relevance." | |
| The experiments conducted by Hall's group are part of a push in their field to expand and generalize the capabilities of solid-state nanopore systems and pursue applications matched to their strengths, including speed of measurement, small required sample sizes, and flexibility in the types and sizes of the analytes that can be probed. | |
| For example, other groups have been applying solid-state nanopores to DNA epigenetics and drug interactions. These pursuits are more likely to yield useful tools and protocols in the near term. | |
| "Going forward, global assessment of depurination levels is an important capability, even more powerful would be the ability to localize the damage to specific genes," Hall says. "Solid-state nanopores may be able to achieve this as well, with their ability to spatially map structural variations at the single-molecule level. We are currently looking into ways to explore this." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
