| Posted: Mar 19, 2015 | |
Fast molecular cargo transport by diffusion |
|
| (Nanowerk Spotlight) Engineered molecular motors are a prime example of nanotechnology's efforts to imitate biology in order to build artificial chemical systems with similar power and capabilities as living cells. For the more visionary goals of nanotechnology, functional and perhaps autonomous molecular motors will play an essential part, just like electric motors can be found in many appliances today. These nanomachines could perform functions similar to the biological molecular motors found in living cells, things like transporting and assembling molecules, or facilitating chemical reactions by pumping protons through membranes (read more: "Off to the races – artificial nanomachines are playing catch-up to nature's molecular motors"). | |
| Researchers have now explored a transport strategy that is different from these nanomotors: They utilized diffusion as an effective transport mechanism for DNA nanotechnology. These findings contribute a new aspect to be considered for the design of future DNA motors, molecular machines, and nanorobots as they provide a simple way to transport molecules over distances of potentially several 100 nm; which is much faster than when using conventional DNA walkers or motors, which make many small and slow steps. | |
| "In contrast to earlier work on DNA walkers, which sought to precisely control the transport path of a DNA walker on a DNA 'track', we give up this precise control and simply 'fly' from the start to the target position," Friedrich C. Simmel, a Professor for Systems Biophysics and Bionanotechnology at the Technische Universität München, tells Nanowerk. "As a benefit, the effective transport speed is much faster." | |
| The team reports their findings in the March 5, 2015 online edition of Nano Letters ("Diffusive Transport of Molecular Cargo Tethered to a DNA Origami Platform"). | |
 |
|
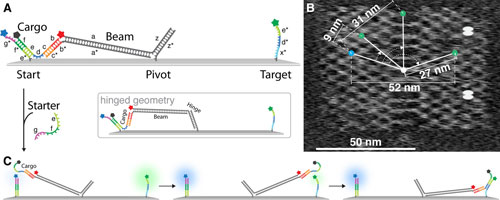
| (A) Schematic side view of a DNA origami platform with a rigid tether fixed in the center. The cargo (domains b, c, d, e, f) can hybridize to sequence domains at the start (e*, f*) and a target (d*, e*) position, as well as to a single stranded extension (b*, c*) of the rigid tether. The inset shows an alternative binding orientation, which results in a more flexible, “hinged” structure. Complementary sequences are indicated by matching colors as well. All sequence domains have a length of 8 nt except for a/a*(88 nt) and z/z*(21 nt). (B) Typical AFM image of the DNA origami rectangle, on which the start position (blue) and three different target positions are indicated. The position of two dumbbell hairpin loops (light gray) is also shown, which are placed asymmetrically to create height contrast for AFM imaging. (C) Mechanism of cargo transport and fluorescent labeling. The cargo DNA is functionalized with a Black Hole Quencher I molecule. Duplex formation with start or target strand reduces the fluorescence intensity of the Atto 488 dye (blue) and Atto 532 dye (green), respectively. An additional Atto 655 dye (red) used as a marker for gel electrophoresis experiments (cf. Supporting Information Figure S2) is permanently attached to the tether and is not affected by quenching. Upon addition of starter strand (domains g, f, e), the beam and cargo are released from the start site, unquenching the blue dye. Upon binding to the target site, the green dye is quenched. (reprinted with permission by American Chemical Society) (click on image to enlarge) | |
| "Our work was motivated by the observation that small biological organisms such as bacteria do not have molecular motors – they are not needed as diffusion within a bacterial cell is very fast anyway," explains Simmel. "Molecular cargo is simply targeted by providing appropriate binding sites for the cargo. As our system is not compartmentalized – such as a bacterial cell – we tether the cargo to our transport platform, which effectively also creates some type of localization or compartmentalization. | |
| He adds that another motivation is a general lesson learned from molecular machines and motors in biology: they utilize ('rectify') Brownian motion rather than work against it. | |
| On the length scale of typical DNA nanostructures (10-100 nm), diffusion is an extremely fast process and thus can be used to transfer a molecule from one position on a nanostructure to another. In order to avoid escape of the cargo it has to be tethered to the DNA structure with a 'transporter arm'. | |
| If DNA molecules are transported (as in this case), they have to unbind from a starting position and later bind to a target position via DNA hybridization i.e., the formation of DNA duplexes between sequence-complementary single strands of DNA. | |
| However, hybridization is a relatively slow process. It turns out that on the scale of the DNA nanostructures used in this work, diffusion is much faster than these binding and unbinding reactions. | |
| "As a consequence, the time required for transport from the starting to the end position is in fact independent from the distance," Simmel points out. "Thus the best – i.e. fastest – strategy to transport DNA molecules in this context is to simply make very large steps." | |
| Using the scaffolded DNA origami technique (see the original 2006 Nature paper "Folding DNA to create nanoscale shapes and patterns"), the team created a rectangular platform with dimensions 90 nm × 60 nm, which was equipped with a 30 nm long molecular tether attached to a flexible hinge in the center of the rectangle. | |
| The tip of the arm can be fixed to an initial position or one of several target positions on the platform, and thus facilitate local transport of molecular cargoes attached to the arm. The role of the tether is to prevent diffusive loss of the cargo, whereas transport itself is driven by thermal motion. | |
| In a typical experiment, the tether is released from its initial position by the addition of a trigger oligonucleotide. The tether then diffusively searches for an alternative binding position on the platform and, finally, attaches to the desired target site. | |
| "Our kinetic studies indicate that this mechanism provides fast transport over distances of up to 52 nm with effective velocities of at least several nanometers per second and, thus, is much faster than transport with DNA walkers," notes Simmel. | |
| Potential applications of this work are found in the field of DNA-templated synthesis: bringing compounds together to increase their local concentration and let them react or interact with each other. In combination with 'mechanical' degrees of freedom, this results in systems that resemble 'molecular assembly lines'. | |
| According to the researchers, another application can be found in DNA computing, where localization of DNA computational gates is supposed to speed up computation processes, reduce crosstalk between competing reactions and thus improve modularity of DNA circuits. | |
| "One could also think of linking many of such systems as ours together and let a molecular signal run through a network of such DNA transporters," says Simmel. "Applications for these types of systems could be found in biosensing, where a biological input signal would be directly and autonomously processed by a biomolecular computation – rather than having to translate it into electrons or photons with a conventional sensor." | |
| Scientists are beginning to learn how to utilize spatial organization and compartmentalization to control the speed and efficiency of (bio)chemical processes, and this work is just one example for that. "In the long run, we hope to create artificial, cell-like systems which have many of the capabilities of biological cells. We are currently exploring components and concepts for such systems," concludes Simmel. | |
| He cautions, though, that one of the biggest challenges for the field is interfacing with the 'outside world' and the combination with other chemistries: "While DNA is a great material to build programmable structures and develop artificial reaction networks, in applications one typically has to deal with other molecules. So the question is whether concepts developed for DNA-based model systems can be translated to other molecular systems, or how to utilize DNA components within hybrid systems." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
