| May 26, 2020 | |
Interfacing synthetic biology with microelectronics |
|
| (Nanowerk Spotlight) Synthetic biology is a field where characterizing genetic circuit behavior is commonly accomplished using fluorescent reporter genes, which can convert a biological response into an optical signal. However, these methods generally require sophisticated instrumentation and may have intrinsic challenges such as the presence of background signals, toxicity, or the need for additional substrates. | |
| Advances in microelectronics have provided a promising alternative in the progress of smart analytical tools, for instance by leveraging the properties of biological molecules in microelectronic sensors. | |
| "Connecting bacterial gene expression to electrodes is an appealing approach to interface genetic circuits with microelectronics for multiple applications," researchers at Prof. Jeff Hasty's lab at UC San Diego's Biocircuits Institute tell Nanowerk. "In particular, we have engineered bacterial circuits capable of controlling the conductivity of the media via cell growth and death. Therefore, the electronic output is controlled by a killing gene. This strategy serves as a general interface between bacteria and electronics compared to previous approaches that relied on electron transfer from bacteria to redox intermediaries." | |
| A paper in Science Advances ("Interfacing gene circuits with microelectronics through engineered population dynamics"), co-first authored by Omar Din and Aída Martín, demonstrates the integration of synthetic biology with electronic circuitry through engineered population dynamics that regulate the accumulation of charged metabolites. The resulting sensor devices can detect changes in bacterial population in response to the presence of chemicals, light or pH. | |
 |
|
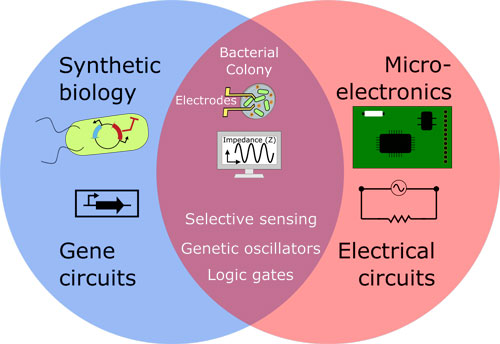
| Interfacing gene circuits with microelectronics through engineered population dynamics. (Image: Aída Martín Galán) | |
| Microorganisms can sense their environment with high selectivity and sensitivity, in some cases better that electronic devices. Designing a new way to 'talk' to bacteria by designing 'bacterial integrated circuits' (BiCs) could lead to a better understanding of the nanoworld for new science advances in synthetic biology, analytical chemistry, and microelectronic technologies. | |
| In analytical chemistry, this work demonstrates opportunities for novel selective electrochemical biosensors, which could be part of new wearable electronics or nanomaterial-modified electrodes. By connecting the bacterial sensing output directly to electrodes, isolation and preparation steps are no longer required to build a biochemical sensor. | |
| In addition, compared to common molecular-based biosensors that use enzymes or antibodies, the proposed lysis triggering of bacteria enables dynamic sensing because cells can be grown in continuous environmental conditions. | |
| "During bacterial growth, charged ions are naturally released because of metabolic processes. As a result, the surrounding medium becomes more conductive, which decreases the impedance to electrical current," Martín explains. "To leverage this phenomenon for the measurement of gene circuit behavior, we engineered the expression of a bacterial killing gene as the output. The resulting behavior can be conceptualized as a bacterial population decreasing in culture impedance over time due to the release of charged ions, after which cell death decreases ion generation and increases impedance." | |
| The researchers monitor electrochemical impedance spectroscopy of the engineered bacterial cultures in milliscale growth chambers that each include electrodes and optical output sensor as a secondary measurement. | |
 |
|
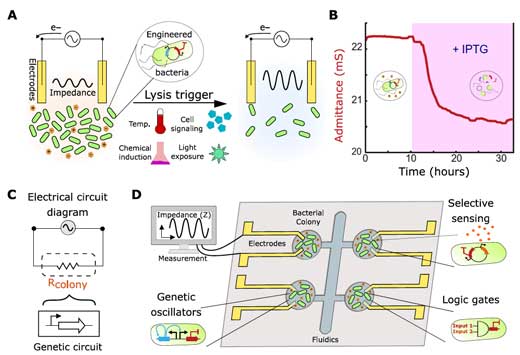
| Interfacing genetic circuits with electrical measurement. (A) Schematic of the approach using a culture of bacteria with a killing gene as the circuit output in contact with inert gold electrodes. The impedance of the culture reduces during growth (left) and increases upon the induction of bacterial death due to the clearance of charged metabolites (right). (B) Profile of the admittance (red), which is the inverse of impedance, using an IPTG-inducible lysis construct (pE35GFP) in an electrochemostat device. The pink shaded region represents induction of lysis with 1 mM IPTG in the medium. (C) Schematic of the equivalent electrical circuit for our strategy using an alternating input voltage. The bacterial population is simplified to a resistor, which is controlled by a genetic circuit. (D) Schematic showing a microelectronic platform to interface between engineered bacteria and electronics. Several chambers may contain unique genetic circuits, connected via electrodes to an impedance output system. (Distributed by AAAS under a Creative Commons Attribution NonCommercial License 4.0 (CC BY-NC)) (click on image to enlarge) | |
| Based on this platform, the team developed a set of applications including a heavy metal sensor via a population control circuit and a dynamic genetic oscillator, which is synchronized to lyse periodically. | |
| According to the team, their design strategy could facilitate advances in synthetic biology, so different genetic circuits (oscillators, logic gates, etc.) may easily be connected to an electrical measure. Also, the use of engineered population control circuits with electrochemical measurement may lead to new opportunities for simple and cost-effective sensing, bypassing optical requirements. | |
| "Three-dimensional layering would also be possible with the use of electrodes compared to conventional imaging methods, as well as in vivo depth tissue measurements using implantable electronics for applications where bacteria reside in vivo," Martín notes. "However, one of the main challenges of our approach is to reduce the response time after triggering lysis since the biosensor relies on the translation of proteins. Thus, further efforts in the analysis and new gene circuits may decrease these times." | |
| Another challenge in measuring bacterial cultures with electrodes is the long-term stability due to biofouling. | |
| Going forward, the researchers will focus on improving the multiplexing of the bacterial platform, which may include isolating the electrodes from the bacterial culture or using microfluidic chambers that exclusively allow the supernatant to be measured. | |
| "We envision that our strategies could lead to the construction of hybrid computational devices, where an array of bacterial colonies process external information through genetic circuitry, which is then transferred via impedance, to an electronic device for analysis," the authors conclude. "The realization of this platform may lead to bICs-based devices that sense environmentally relevant molecules or enable unexplored investigations of engineered bacterial populations in the context of synthetic biology." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
