| Posted: Jun 22, 2006 | |
A milestone towards fast DNA sequencing with nanopore devices |
|
| (Nanowerk Spotlight) Setting a milestone towards fast DNA sequencing by a nanopore device, researchers developed a solid state nanopore device that can detect the difference between single molecules of double and single stranded DNA at high speed, with high accuracy, and at extreme pH. This research is a key step to develop a nanopore sequencing machine. More immediate application may be the sensing of long chain polymers for medical diagnostics. | |
| Nanopores function as membrane channels in all living systems, where they serve as sensitive electro-mechanical devices that regulate electrical potential, ionic flow, and molecular transport through the cell membrane. Scientists are studying nanopore construction with the goal of making man-made cell membranes and single molecule detectors. | |
| For a nanopore to be useful as a single molecule detector, its diameter must not be much larger than the size of the molecule to be detected. When a single molecule enters a nanopore in an insulating membrane, it causes changes in the nanopore's electrical properties, which then can be detected and measured. A high-throughput solid state nanopore device that can probe and directly "read" electronically, at the single molecule level, the size, folding, and eventually the sequence of DNA and proteins, will dramatically alter the pace of biology and medical science. | |
| "Single molecule methods based on nanopore detectors provide a new approach to rapid characterization and perhaps even sequencing of long biomolecules such as DNA" Professor Jiali Li explains to Nanowerk. Li, together with her colleagues at the University of Arkansas and collaborators at the Department of Physics at Harvard University, led by Professor Jene A. Golovchenko, published a paper ("Detecting Single Stranded DNA with a Solid State Nanopore") on her findings in Nano Letters. | |
| "Our work is the first demonstration of "raw" denatured DNA detected by any (biological or solid state) nanopore device and the first observation any single stranded DNA translocating freely through a solid state nanopore device" Li points out. "It allows us to determine the distribution of a mixture of double and single stranded DNA. Single molecule identification of polymers by diameter and length are now possible." | |
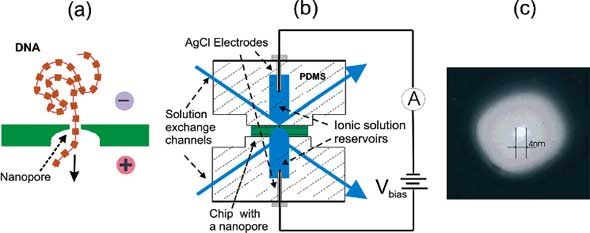 | |
| (a) Schematic illustration of DNA molecule translocating through a solid-state nanopore. (b) Experimental setup for single molecule measurements with a nanopore detector. (c) TEM of a silicon nitride nanopore, in this case 4 nm in diameter in a ∼5-10 nm thick local membrane. (Reprinted with permission from the American Chemical Society) | |
| Solid-state nanopores are eminently suited for DNA and protein detection because they are mechanically robust, have tunable dimensions, and tolerate broad temperatures, pH, and chemical variations. | |
| The nanopores for this research were made in silicon nitride (Si3N4) as an insulating membrane, focused ion beam milling, followed by feedback controlled ion beam sculpting. The experimental setup consists of an ionic solution divided into two isolated reservoirs by the membrane containing a single nanopore. An ionic current through the open nanopore is established by a voltage applied between silver/silver chloride electrodes placed in each of the two reservoirs. DNA molecules are added to the reservoir with the negative electrode. The electric field within the nanopore then induces the molecule to pass through it to the positive reservoir. The resulting changes of the nanopore current are then measured and document the DNA passing through it. | |
| "Our experiment demonstrates conclusively that long denatured single stranded DNA molecules can be observed to freely translocate in stretched out configurations through a solid-state nanopore" Li says. "We are now working on increasing the spatial and temporal resolution of the nanopore detecting system as the next important goal." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
