| Aug 04, 2014 |
How the 'biological spark plug' in biomolecular motors works
|
|
(Nanowerk News) Using high-performance computers and quantum mechanical methods, researchers at Heidelberg University have simulated processes that reveal how the “biological spark plug” works in the biomolecular motors of cells. Under the direction of Dr. Stefan Fischer, the investigations focused on the myosin protein, which, among other things, is responsible for muscle movement. The researchers’ extensive simulations show how the release of energy is initiated in this complex motor.
|
|
The results of the research conducted at the Interdisciplinary Center for Scientific Computing were published in the journal PNAS ("Catalytic strategy used by the myosin motor to hydrolyze ATP").
|
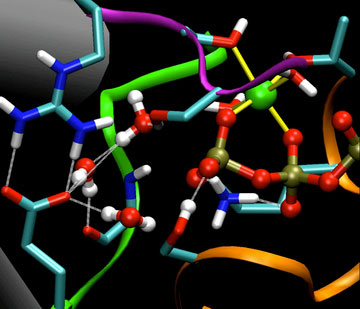 |
| ATP binding pocket of myosin. The illustration shows a portion of the atoms that have to move during the catalytic division of ATP.
|
|
Biomolecular motors are protein molecules responsible for mechanical movement in cells. These smallest of known motors use the molecule adenosine triphosphate (ATP) as fuel, which all living organisms use as a source of energy for processes that require it.
|
|
In order to understand how these cell motors use ATP to function, they can be compared to an automobile engine, in which energy is released by burning petrol. Because petrol does not ignite by itself, energy must be applied to initiate the combustion reaction. This job is done by the spark plug. Energy is not released until the heat energy of the spark is applied to overcome the energy barrier of petrol combustion. According to Stefan Fischer, there are a number of parallels to biomolecular motors. The ATP molecule is stable and like petrol does not release its energy spontaneously. Whereas ATP splits rather than burns, there is also an energy barrier that must be crossed to trigger that splitting, known as hydrolysis.
|
|
Dr. Fischer’s research team studied exactly how the trigger mechanism for energy release works in biomolecular motors. “We wanted to find out how the energy stored in the ATP gets released so selectively and precisely timed,” explains the Heidelberg researcher, who heads the Biological Macromolecules working group at the Interdisciplinary Center for Scientific Computing (IWR).
|
|
The scientists launched their study of the “biological spark plug” using the biomolecular motor myosin. Myosin is a family of motor proteins that uses ATP, for example to drive muscle movement. The ATP is bound in a sort of “pocket” in the protein. The pocket lowers the energy barrier for splitting the ATP – this process of lowering is known as catalysis – and ensures that the desired chemical reaction ensues and ultimately energy is released. The “catalytic pocket” is the biological equivalent of the spark plug in the combustion engine, according to Dr. Fischer.
|
|
The existence of this “biological spark plug” has been known for more than 50 years, but researchers have never been able to fully explain how it works, as Stefan Fischer emphasises: “The reaction takes place in about a trillionth of a second, pushing experimental methods to their limits. This event could not be studied exactly until the computer-assisted methods of scientific computing were applied.” The scientists first had to identify which of the 6,000 atoms of myosin were essential for catalysis. After comprehensive simulations lasting several years, the researchers identified the role of approximately 200 relevant atoms. Because both the myosin atoms and ATP atoms must move during ATP hydrolysis, the possibilities for movement in three-dimensional space are countless – though only one path leads to the lowest energy barrier. “We had to calculate the paths of all approximately 200 atoms in three dimensions; altogether a problem in 600 dimensions,” says Dr. Fischer.
|
|
For their complex calculations, the scientists combined the scientific methods from quantum mechanics with high-performance computers. This allowed them to clarify how the interactions between ATP and myosin are organised in order to lower the energy barrier for splitting ATP. Stefan Fischer explains that the electrostatic charges on the protein atoms are positioned around the ATP in such a way that they modify the electron density of this molecule, making it easier for the ATP fuel to split. “This way we could precisely quantify how much every myosin atom relevant in this process contributed to lowering the energy barrier. Based on these findings we succeeded in clearly formulating the protein’s catalytic strategy.”
|
|
The biological spark plug mechanism described by the IWR researchers is not only found in cell motors, but is probably also used in all other protein molecules that use ATP as an energy source, says Dr. Fischer. “Because ATP is the fundamental energy currency of cells, almost all biochemical processes in the body are concerned. In terms of a practical application, our findings may be able to help research on new medications for treating cardiac muscle diseases. Our discoveries may also spur new approaches to treating diseases in which ATP splitting is a part of the biochemistry of the pathological system.”
|

