| Posted: Jan 10, 2007 | |
Nanorobotic arm to operate within DNA sequence |
|
| (Nanowerk Spotlight) The success of nanorobotics requires the precise placement and subsequent operation of specific nanomechanical devices at particular locations, thereby leading to a diversity of structural states. The structural programmability of DNA makes it a particularly attractive system for nanorobotics. A large number of DNA-based nanomechanical devices have been described, controlled by a variety of methods. These include pH changes and the addition of other molecular components, such as small molecule effectors, proteins and DNA strands. The most versatile of these devices are those that are controlled by DNA strands. This versatility results because they can be addressed specifically by strands with particular sequences. Researchers at New York University have developed a framework that contains a binding site – a cassette – that allows insertion of a rotary device into a specific site of a DNA array, allowing for the motion of a nanorobotic arm. Changing the cassette’s control sequences or insertion sequences allows the researchers to manipulate the array or insert it at different locations. | |
| "We have developed a cassette that enables us to direct a device to a particular place in an array" Dr. Nadrian C. Seeman, professor at NYU's Chemistry Department, explains to Nanowerk. "This is the first time that a sequence-dependent DNA-based nanomechanical device has been targeted to a specific site in a 2D lattice. This means that ultimately a variety of devices which can be in multiple states, and which are individually addressable, can be organized in lattices, with a large number of states associated with them. The two states of this device can be distinguished by atomic force microscopy." | |
| The insertion of such nanomechanical devices into 2D arrays results in a nanorobotic system, wherein nanoscale moving parts can be controlled relative to a fixed frame of reference. | |
| Seeman points out that the results pave the way for creating nanoscale 'assembly lines' in which more complex maneuvers could be executed. | |
 |
|
| (click on image to enlarge) Schematic depiction of the cassette. (A) A view perpendicular to the plane of the cassette in the PX state. The PX state is set by the green strands in the middle of the upper two domains. The reporter hairpin is seen end-on protruding from the plane. The sticky ends on the bottom domain attach the cassette to the 2D array. (B) The same molecule is shown obliquely so the reporter hairpin can be seen. (C) A view similar to (A), except that the cassette is in the JX2 state, which is set by the purple strands. The reporter hairpin is now behind the cassette, a point emphasized in (D). All drawings are in a virtual-bond representation produced by the program GIDEON. (E) A 5% polyacrylamide gel run in TAEMg (Tris Acetate EDTA Magnesium) buffer . The two different states are shown both for a cassette without a hairpin and for a cassette including a reporter hairpin. The single bands in each lane indicate that the motifs are stable and monodisperse. BP, base pair. (Graphic: Dr. Seeman) | |
| The results are based upon a device Seeman and (then) NYU graduate student Hao Yan had previously developed (Yan is now a professor at Arizona State University). That component has enabled the translation of DNA sequences, thereby potentially serving as a factory for assembling the building blocks of new materials. The invention has the potential to develop new synthetic fibers, advance the encryption of information, and improve DNA-based computation. | |
| Another previous device, developed with NYU Chemistry graduate student Shiping Liao, contains two PX-JX2 devices and emulates the process by which RNA replicas of DNA sequences are translated to create protein sequences. However, the signals that control the nanomechanical tool are DNA rather than RNA. The dimensions of the machine are approximately 110 x 30 x 2 nm. | |
 |
|
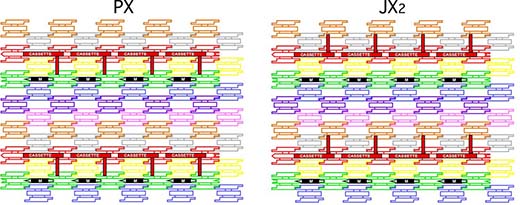
| (click on image to enlarge) The arrays are shown schematically to demonstrate the two states of the device in the cassette. The eight TX tiles that form the array are shown in differently colored outlined tiles. For clarity, the cohesive ends are shown to be the same geometrical shape, although they all contain different sequences. The cassette and reporter helix are shown as solid red components; the marker tile is labeled M and is shown with a solid black rectangle representing the domain of the tile that protrudes from the rest of the array. Both the cassette and the marker tile are rotated ∼103° from the other components of the array (three nucleotides rotation). The PX arrangement is shown on the left, and the JX2 arrangement is on the right. The reporter hairpin points toward the marker tile in the PX state but points away from it in the JX2 state. (Graphic: Dr. Seeman) | |
| "It is crucial for nanorobotics to be able to insert controllable devices into a substrate, thereby leading to a diversity of structural states" says Seeman. "Here we have demonstrated that a single device has been inserted and converted at a specific site. There is no reason to expect that the system is limited to a single device unit." | |
| Seeman's latest findings have been reported in Science "Operation of a DNA Robot Arm Inserted into a 2D DNA Crystalline Substrate". | |
| Also see our previous Nanowerk Spotlight that highlights the control of the PX-JX2 device by RNA. This method was used in the above work as well. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
