| Posted: Jan 12, 2010 | |
Simple DNA nanomachine is capable of continuous rotation |
|
| (Nanowerk Spotlight) Sophisticated molecular-size motors have evolved in nature, where they are used in virtually every important biological process. Scientists around the world are working hard to develop synthetic nanomotors that mimic the function of these amazing natural systems and could be used in man-made nanodevices. A tremendous complication for nanotechnology researchers is the fact that building nanoscale motors is not just an exercise in scaling down the design of a macroworld engine to nanoscale dimensions. Many factors such as friction, heat dissipation and many other mechanical behaviors are just very different at this scale – everything is constantly moving (under kinetic energy supplied by the heat of the surroundings) and being buffeted by other atoms and molecules (Brownian motion). | |
| A very promising field of nanomotor research are DNA nanomachines (see "Synthetic DNA nanomachines go to work inside living cells"). These are synthetic DNA assemblies that switch between defined molecular shapes upon stimulation by external triggers. They can be controlled by a variety of methods that include pH changes and the addition of other molecular components, such as small molecule effectors, proteins and DNA strands. | |
| Researchers have now designed and built a simple DNA machine that is capable of continuous rotation with controlled speed and direction – a function that might be very useful for example for molecular transport. | |
| "Our machine is driven by an externally controlled electric field" Daniel Lubrich tells Nanowerk. "When this field is oscillated between four directions, it continuously reorients a rotor DNA that is asymmetrically attached to a DNA axle. The axle is held in place between a glass surface and a magnetic bead. Single-stranded segments on the DNA axle act as bearings, providing for free rotation around the molecular bonds between nucleotide bases. This design hence allows for free and continuous rotation of the rotor DNA around the axle as it reorients with the oscillating electric field." | |
| Lubrich, a Research Fellow at National University of Singapore's NanoCore, and his collaborators from Forschungszentrum Karlsruhe in Germany and Harvard University, have presented their work in a recent online edition of Small ("A Rotational DNA Nanomotor Driven by an Externally Controlled Electric Field"). | |
 |
|
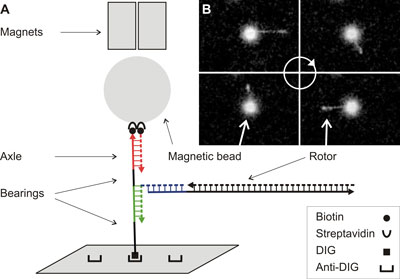
| Design of the rotational DNA nanomachine. A) Schematic representation of the design of the rotational DNA machine as well as its suspension between a functionalized glass cover slip and a magnetic bead. DNA and DNA oligonucleotides are drawn as lines. The black DNA sequences on the axle do not have a complement (they are single-stranded) and act as bearings, allowing for free rotation of the rotor around the axle. As is customary, the arrows on the DNA point towards the 3' end. The magnetic bead is held fast by a pair of magnets. The scheme is not drawn to scale. B) Snapshots from a video taken in a microscope. In the microscope the rotational DNA nanomachine is viewed from the top. Shown are four different orientations of the rotor DNA. Reorientation of the rotor with the oscillating electric field leads to continuous rotation of the rotor around the axle. (Image: Daniel Lubrich, National University of Singapore) | |
| To construct their DNA nanomotor, the team uses as axle a 30nm-long DNA oligonucleotide, modified at both ends. The 5' end is modified with digoxigenin NHS ester (DIG); the 3' end is modified with biotin. This allows for simultaneous attachment to an anti-DIG modified glass surface and a steptavidin modified magnetic bead. Lubrich notes that this sandwiches the axle and keeps it firmly in place when forces between magnets and the magnetic bead pull the axle tight. The short DNA oligonucleotides which are drawn as colored lines in the figure above act as 'nuts and bolts' for attaching the various parts of the system to each other. | |
| "The rotor DNA used in our work is λ-DNA with an approximate contour length of 16µm. λ-DNA was used because it is cheap and because it is easy to see in the microscope. Shorter DNA can however also be used. Observation of the rotation in the microscope will then be a little bit harder. The rotor DNA, like all DNA at ambient pH, is negatively charged" explains Lubrich. "It is bound to the axle oligo in an asymmetric fashion but is otherwise unrestricted. Its role is to align with an electric field. Switching the orientation of this field will hence cause the rotor DNA to change its orientation relative to the axle. Since the rotor is attached to the axle in between two short single-stranded sections on the DNA axle, acting as bearings by allowing for free rotation around molecular bonds between nucleotide bases, this reorientation leads to rotation of the rotor DNA around the axle DNA. The axle is firmly held in place, sandwiched between a glass surface and a magnetic bead, which is held fast by external magnets and is hence acting as stator." | |
| The speed and direction of the rotational motion can be controlled by adjusting the oscillation frequency and direction of the electric field. While Lubrich and the team have concentrated on continuous rotation, they point out that any motion that is within the intrinsic speed limit of the rotational DNA nanomotor – and starts and ends at one of the four possible orientations of the rotor DNA – is possible in principle. | |
| Rotational DNA nanomotors may find applications as nanofluidic pumps, for transport of cargo, nanoscale stirring and mixing, as well as the forcing of mechanical interactions between the DNA rotor (or anything that is attached to it) and obstacles that could be placed in its way. | |
| According to Lubrich, future directions and challenges in the field are related to the development of much better nanodevices. "They should run as autonomous as possible and be programmable as to perform certain functions, start on command, pause, stop, restart, go backwards, forwards, and so on. It is probably also important to work towards having a 'toolkit' of standard parts which can be used and re-used in a variety of different settings. This would then enable nanoscientists and -engineers to build nanodevices quicker and better, without having to invent the whole thing all the time." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
