| Posted: Feb 07, 2013 | |
DNA nanoarchitectonics |
|
| (Nanowerk Spotlight) DNA is a powerful biomaterial for creating rationally designed and functionally enhanced nanostructures. Emerging DNA nanotechnology employs DNA as a programmable building material for self-assembled, nanoscale structures (read more: "Reality check for DNA nanotechnology"). Researchers have also shown that DNA nanotechnology can be integrated with traditional silicon processing. DNA nanoarchitectures positioned at substrate interfaces can offer unique advantages leading to improved surface properties relevant to biosensing (for instance, graphene and DNA can combine to create a stable and accurate biosensor), nanotechnology, materials science, and cell biology. | |
| A new Perspectives article in Langmuir ("DNA Nanoarchitectonics: Assembled DNA at Interfaces") by Stefan Howorka, an associate professor in the Department of Chemistry at University College London, highlights the benefits and challenges of using assembled DNA as a nanoscale building block for interfacial layers. | |
| After assessing how assembled DNA compares to more conventional polymeric building blocks, Howorka discusses three areas of applications of DNA nanoarchitectonics: homogeneous films with an emphasis on biomolecular recognition; 2D nanopatterns that are produced via the bottom-up route in optional combination with top-down nanofabrication; and 3D superlattices of nanoparticles. In a concluding section he identifies some possible areas of future research for interfacial DNA nanostructures. | |
| Thin-films and surface coatings | |
| Efficient and correct binding of biomolecules is relevant in biosensing, purification, and biophysics. However, the specific interaction of molecules at interfaces can be impaired by restricted target accessibility caused by imperfectly packed receptors on locally crowded surfaces. | |
| Homogeneous films made up of DNA nanoarchitectures were able to overcome some of these problems and molecularly thin and laterally dense films composed of DNA structures have been constructed to improve the biomolecular recognition at biointerfaces. | |
| Bottom-up nanopatterning | |
| In addition to the homogeneous coatings of substrates, DNA nanoarchitectures are predominately utilized to pattern surfaces on the nanoscale. In principle, nanopatterning with DNA can be achieved either in a bottom-up fashion or in combination with top-down approaches. In both cases, producing nanopatterns can open up many applications in nanotechnology, biophysics, and cell biology. | |
| Bottom-up nanopatterning is done with DNA tiles (see: "Scientists develop nanodevice manufacturing strategy using DNA building blocks") that can be programmed to assemble themselves into precisely designed shapes, such as letters and emoticons. Further development of the technology could enable the creation of new nanoscale devices, such as those that deliver drugs directly to disease sites. | |
| It can also be done with DNA origami, a technique that addresses some of the limitations of DNA tiles. DNA origami is a method for folding long strands of DNA into whatever very small shape or pattern you desire. Using a computer-aided design program a scientist can design the desired nanoscale shape (typically about 100 nanometers across) and the computer designs a set of short DNA strands that can be ordered from a company that specializes in synthesizing DNA strands. The short DNA strands get mixed with the long DNA strands, heated up to nearly boiling, and cooled to room temperature over the course of a couple hours. In a single drop of water one then has 100 billion copies of the desired shape, or shape with a pattern on top. In the first DNA origami made were shapes like triangles and smiley faces, and patterns like maps of the western hemisphere, snowflakes, etc. | |
| DNA origami is also versatile with regard to the substrates onto which they can bind, as shown for gold, silicon, and graphene. | |
 |
|
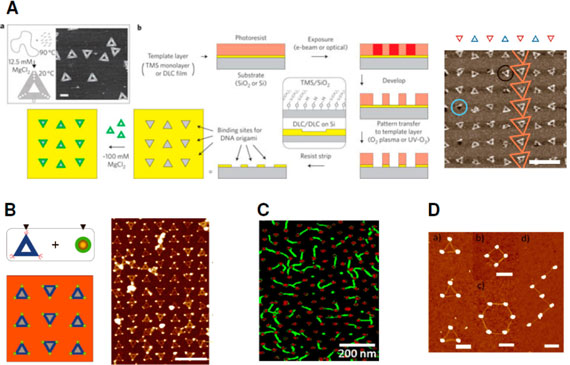
| Nanopatterns generated by combining bottom-up DNA nanoarchitectures and physical top-down routes. (A) Generation of patterns of DNA triangles. (a) Synthesis scheme for DNA origami triangles and atomic force microscopy height image showing random deposition on mica. Scale bar, 100 nm. (b) Fabrication of DNA origami binding sites via photolithography. The inset highlights the differentiation of the background and features (background/features) for the trimethylsilyl (TMS) monolayer and diamondlike carbon (DLC) films. Silanol groups occur in oxidized areas of the TMS monolayers. Features etched into the DLC template layer are 0.5-1.5 nm deep. DNA triangles can bind into the templated layer featuring complementary-shaped binding sites as illustrated by the AFM height image (right) of DNA triangles on 110 nm patterned triangle sites on a DLC/DLC surfaces. (B) Two-dimensional nanoparticle arrays are formed by binding DNA-coated nanocrystals (green with a golden core) onto the edges of DNA origami triangles via hybridization. The hybrid structures adsorb onto the electron-beam lithographically nanopatterned surface that features hydrophilic triangular binding sites (gray) surrounded by a hydrophobic coating (orange). AFM image of gold-nanoparticlefunctionalized origami triangles bound to top-down generated columns of equilateral triangles with sides of 100 nm, alternately oriented up and down. Scale bar, 500 nm. (C) Block-copolymer-patterned arrays of 5 nm gold nanoparticles (red) were connected with DNA origami (green). The connection was mediated by hybridization between the single-stranded DNA-thiol-modified gold nanoparticles and the sticky end-modified DNA origami. (D) Structures formed by connecting lithographically generated gold islands with DNA origami tubes: (a) triangle, (b) hexagon, (c) square, and (d) z shape. All scale bars are 300 nm. The DNA origami nanotubes of approximately 380 nm in length were composed of six DNA helices bundled with a hexagonal cross section and carried thiolated groups near their termini. The distance between the thiolated groups located near each end is approximately 320 nm. (Reprinted with permission from American Chemical Society) | |
| The alternative route to nanopatterned surfaces combines bottom-up DNA materials with a top-down nanofabrication approach. This combination is highly synergistic because the separate strategies cover two different length scales. Whereas the physical top-down approach can pattern microscale areas with feature sizes frequently down to 10-100 nm, the bottom-up approach can afford atomically precise DNA nanostructures of up to but not beyond overall dimensions of a few hundred nanometers. | |
| 3D nanoparticle lattices | |
| Howorka writes that another exciting area of DNA nanoarchitectonics is concerned with colloids and suspensions of nanoparticles. The nanobio interface at spherical substrates is of great interest because it provides several unique advantages and characteristics not available at flat substrates, such as the ability to self-assemble nanoparticles into complex 3D supramolecular arrays. | |
| In concluding his article, Howorka points to two growth areas for DNA nanoarchitectonics: One area is the increased exploitation of DNA origami for 3D lattices of nanoparticles. In the future, this might lead to the creation of photonic crystals with switchable collective properties that are controlled by DNA interparticle spacers of tunable distance. In more general terms, DNA nanostructures at interfaces could be integrated into functional devices whose properties can be actuated by external triggers. | |
| He points out that an increasing trend in DNA nanotechnology is a focus on applications that ultimately lead to commercially viable products. "Although an emphasis on real-world applications is understandable, one should also consider that DNA nanoarchitectonics at interfaces is a relatively new research area compared to that of very established polymeric film coatings. Nevertheless, research activities are underway toward high added-value devices where the higher costs of DNA scaffolds are justified. In conclusion, DNA architectonics at interfaces is a highly interdisciplinary field that spans the nano- and microscale with numerous applications in materials and life science." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
