| Posted: May 06, 2015 | |
Snail-inspired nanosensor detects and maps mRNA in living cells |
|
| (Nanowerk Spotlight) Every cell is a busy messaging center, with thousands of genes churning out messenger RNA (mRNA) transcripts for translation into functional proteins. Cell fate, function, and phenotype are significantly dictated through the spatiotemporal control of mRNA expression. Therefore, determining the mRNA expression of an individual cell can reveal critical insights into that particular cell's health and physiological state. | |
| "Presently, several techniques for detecting mRNAs are available, which include in situ hybridization and polymerase chain reaction (PCR)," says David T. Leong, an Assistant Professor in the Department of Chemical & Biomolecular Engineering at National University of Singapore (NUS). "However, these single-point and end-point techniques require the killing of the cells and are thus unable to capture the expression of mRNA in real time and locality with high precision." | |
| In new work, Leong and his team describe a new way of preparing functional DNA nanostructures that can provide accurate quantification and visualization of mRNA transcripts in living cells. | |
| They reported their findings in the April 23, 2015 online edition of ACS Nano ("Nature-Inspired DNA Nanosensor for Real-Time in Situ Detection of mRNA in Living Cells"). | |
| "One key feature of our DNA-based mRNA sensor design is that it is highly resistant to non-specific enzymatic degradation, and thus was shown to significantly reduce the likelihood of generating false-positive signals," Leong explains to Nanowerk. "This is made possible because of our snail-inspired design. Just like how the shell of the snail protects the snail from the harsh environment but does not completely restrict it from sensing its surroundings, we incorporated this 'protective-yet-accessible' design concept into our DNA nanodevice which we termed nano-SNail-inspirEd Locator (Nano-SNEL)." | |
 |
|
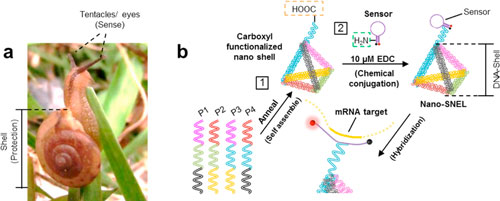
| Design and working principle of nano-SNEL. (a) Image of the Asian tramp snail (Bradybaena similaris) that shows the typical anatomy of the gastropoda species, comprising a shell that is meant to provide essential protection from the harsh environment and whose movement is guided by the sensory organs. (b) Two-step synthesis scheme employed to construct nano-SNEL. The nanosensor (molecular beacon – MB) is chemically conjugated to a pyramidal-shaped nanoshell that was designed to facilitate cellular entry and protect the sensor from nondiscriminatory digestion, which could lead to false negatives. (Reprinted with permission by American Chemical Society) (click on image to enlarge) | |
| Nano-SNEL has two main components: A sensor/reporter to detect mRNA; and a DNA-pyramid shell. The DNA pyramid shell not only serves as a carrier for the sensor to enter easily into a cell but also functions as a decoy or sacrificial shell to protect the sensor from enzymatic attack. | |
| "We showed that our nano-SNEL could synergistically present itself as an excellent intracellular probe, without interfering with either specific targeting by or hybridization-induced fluorescence of the probes," says Leong. "When the cells were treated with nano-SNEL, we were able to attain robust in situ hybridized signals and the associated intracellular spatial distribution could be mapped since the cell itself is not destroyed." | |
| A key feature of nano-SNEL is that it is very stable even under harsh physiological conditions. This opens up a myriad of possible applications where DNA based nanomaterials can be applied directly. | |
| Because the targets – DNA, mRNA, miRNA – are ubiquitous in all biological systems, nano-SNEL is highly versatile as a diagnostic tool. For example, defective expression of mRNA linked to certain heterogeneous diseases such as cancer might be detectable at the single cell resolution using nano-SNEL. | |
| Outside of the cell, Nano-SNEL can also be used to detect viral RNA that may be found in blood serum such as the Ebola virus. This is not possible with mRNA sensors that are susceptible to non-specific enzymatic degradation. | |
| While there are ground breaking and important reports focusing on fabricating exquisite and complex 3D DNA nanostructures, the NUS team stood on the shoulders of the 'DNA nanostructures designer' giants and focussed on applying their fascinating science to problems. This work represents their contributions to the field as a small step towards closing the gap between basic and applied DNA nanosciences. | |
| "In our lab, we are particularly interested in the use of DNA based nanomaterials for biomedical applications," notes Leong. "Compared to other types of classical organic/inorganic nanomaterials, DNA as a malleable putty for nanomedicine is still at its infancy and thus leaves some uncharted territory for exploration. We hope to share with the community a few interesting applications arising from the use of this exciting starting material." | |
| "The use of DNA based materials for nanomedicine applications has gained significant traction in recent years; we believe future studies will need to establish a better understanding of the critical DNA nanostructures-cell biology relationship," concludes Leong. "With these critical insights, we hope to develop smarter and more sophisticated DNA nano-devices to meet some of the challenges of modern medicine. We cannot do all these on our own as the success of DNA nanomaterials as a functional material for biomedical applications definitely requires the cross pollination of ideas from colleagues of diverse backgrounds and expertise." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
